Creates a phylo object from a taxonomy table with hierarchical taxonomic ranks as columns.
Usage
taxo2tree(
physeq,
ranks = c("Domain", "Phylum", "Class", "Order", "Family", "Genus", "Species"),
internal_node_singletons = TRUE,
use_taxa_names = TRUE
)Arguments
- physeq
(required) A
phyloseq-classobject- ranks
Character vector specifying the column names to use as taxonomic ranks, ordered from highest to lowest. By default: c("Domain", "Phylum", "Class", "Order", "Family", "Genus", "Species").
- internal_node_singletons
Logical, if TRUE, create internal nodes for singleton. If FALSE, internal nodes with only one descendant are discarded.
- use_taxa_names
Logical, if TRUE (default), use the taxa names (rownames, e.g., ASV_1, ASV_2) as terminal leaves. If FALSE, collapse identical taxonomy paths and use the lowest rank value as tip labels. This is useful for creating cleaner trees that show only taxonomic structure without individual ASV/OTU names.
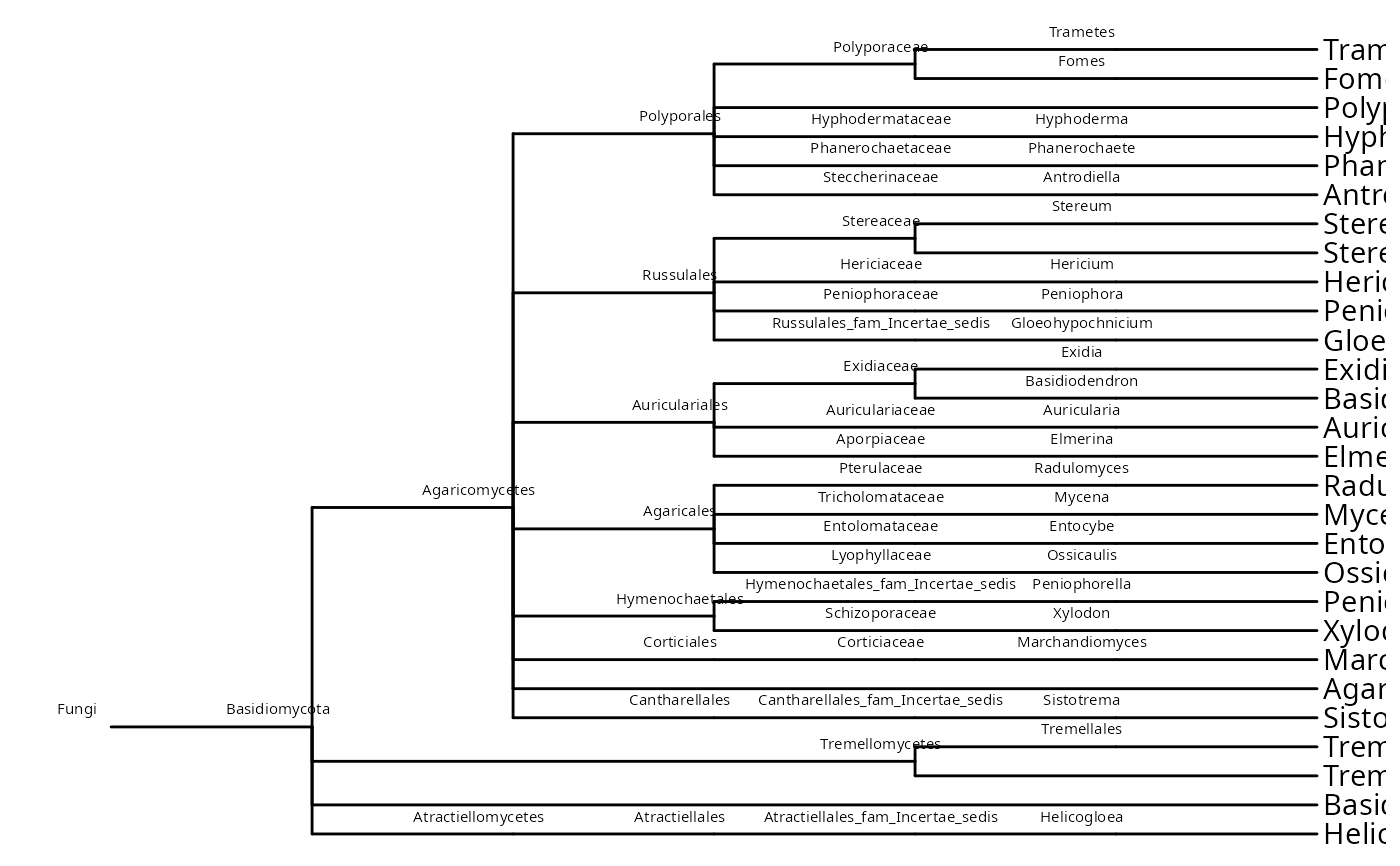
Examples
data_fungi_mini@phy_tree <-
phyloseq::phy_tree(taxo2tree(data_fungi_mini,
ranks = c(
"Domain", "Phylum", "Class", "Order",
"Family", "Genus", "Genus_species"
)
))
#> Found more than one class "phylo" in cache; using the first, from namespace 'phyloseq'
#> Also defined by ‘tidytree’
library(ggtree)
#> ggtree v4.0.4 Learn more at https://yulab-smu.top/contribution-tree-data/
#>
#> Please cite:
#>
#> S Xu, Z Dai, P Guo, X Fu, S Liu, L Zhou, W Tang, T Feng, M Chen, L
#> Zhan, T Wu, E Hu, Y Jiang, X Bo, G Yu. ggtreeExtra: Compact
#> visualization of richly annotated phylogenetic data. Molecular Biology
#> and Evolution. 2021, 38(9):4039-4042. doi: 10.1093/molbev/msab166
#>
#> Attaching package: ‘ggtree’
#> The following object is masked from ‘package:MiscMetabar’:
#>
#> multiplot
ggtree(data_fungi_mini@phy_tree) +
geom_nodelab(size = 2, nudge_x = -0.2, nudge_y = 0.6) +
geom_tiplab()
#> Found more than one class "phylo" in cache; using the first, from namespace 'phyloseq'
#> Also defined by ‘tidytree’
 psm <- psmelt(data_fungi_mini) |>
group_by(OTU) |>
summarize(
Mol_Abund = sum(Abundance),
Frequency = sum(Abundance > 0),
Class = tidyr::replace_na(unique(Class), "Unknown")
)
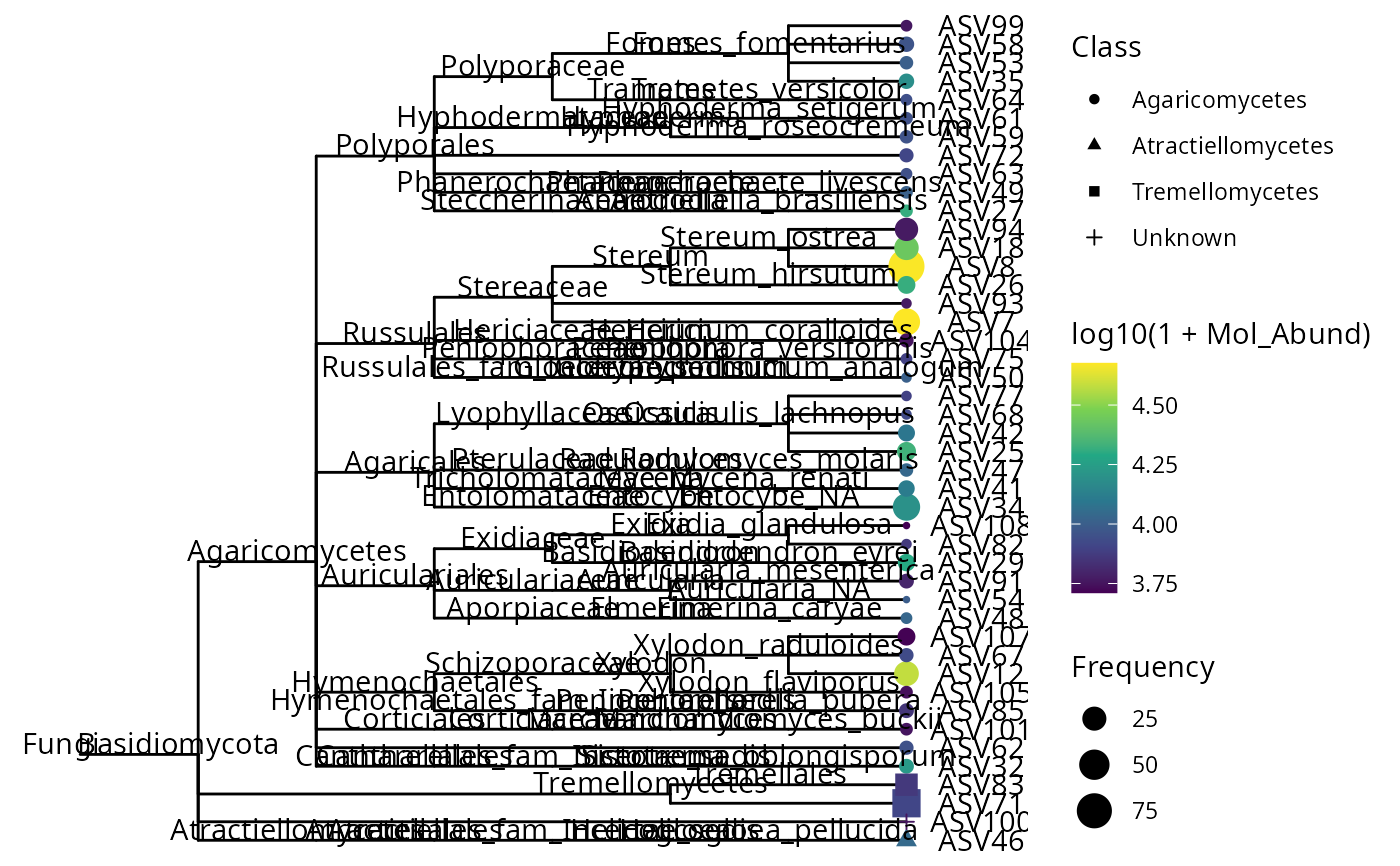
ggtree(data_fungi_mini@phy_tree) %<+% psm +
geom_tiplab(offset = .6, hjust = .5) +
geom_tippoint(aes(
shape = Class,
color = log10(1 + Mol_Abund),
size = Frequency
)) +
geom_nodelab(nudge_x = -0.2, nudge_y = 0.6) +
scale_color_viridis_c()
#> Found more than one class "phylo" in cache; using the first, from namespace 'phyloseq'
#> Also defined by ‘tidytree’
psm <- psmelt(data_fungi_mini) |>
group_by(OTU) |>
summarize(
Mol_Abund = sum(Abundance),
Frequency = sum(Abundance > 0),
Class = tidyr::replace_na(unique(Class), "Unknown")
)
ggtree(data_fungi_mini@phy_tree) %<+% psm +
geom_tiplab(offset = .6, hjust = .5) +
geom_tippoint(aes(
shape = Class,
color = log10(1 + Mol_Abund),
size = Frequency
)) +
geom_nodelab(nudge_x = -0.2, nudge_y = 0.6) +
scale_color_viridis_c()
#> Found more than one class "phylo" in cache; using the first, from namespace 'phyloseq'
#> Also defined by ‘tidytree’
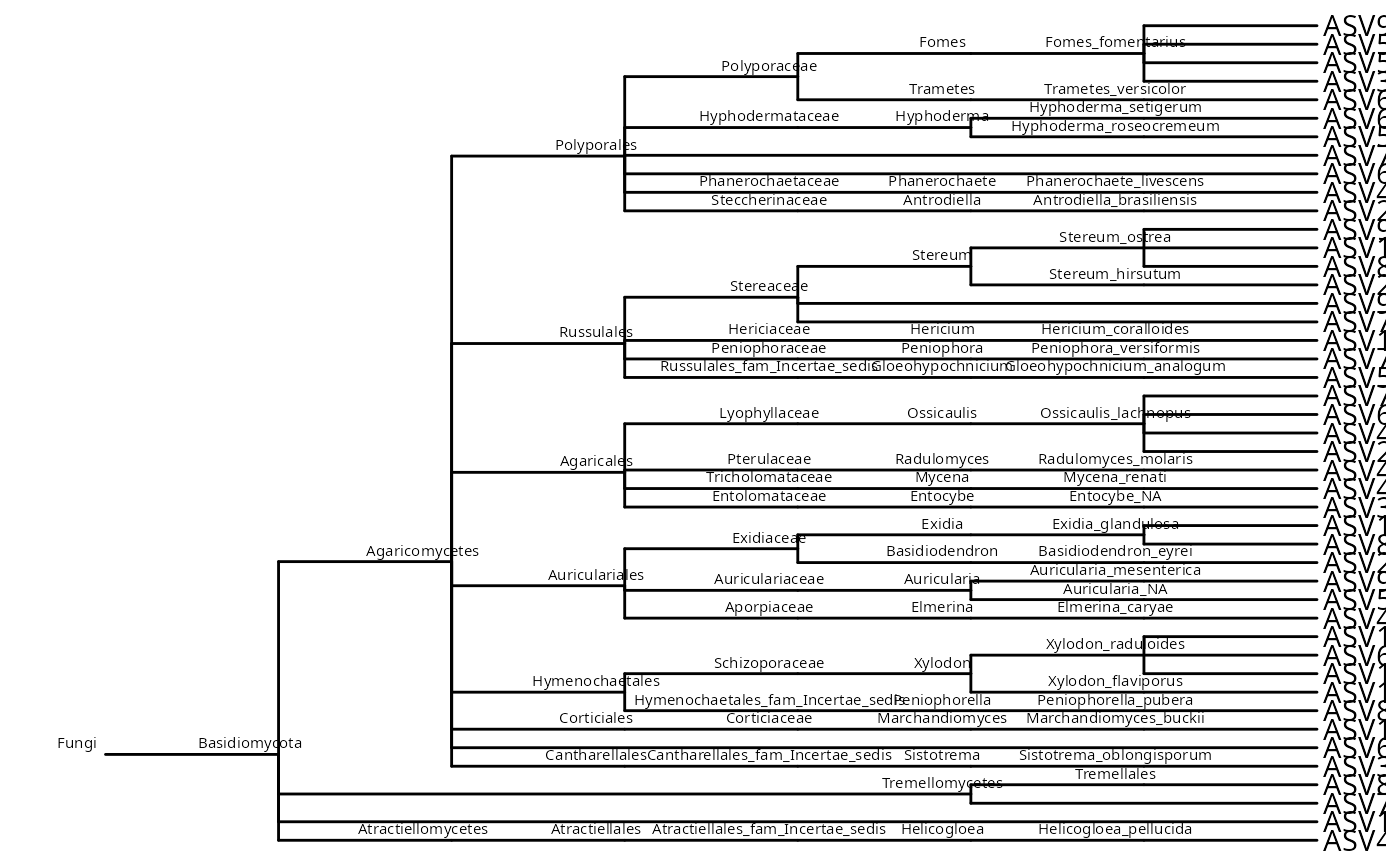
 # Without internal node singletons and only until the Genus level
tree_wo_singletons <-
phyloseq::phy_tree(taxo2tree(data_fungi_mini,
ranks = c(
"Domain", "Phylum", "Class", "Order",
"Family", "Genus"
),
internal_node_singletons = FALSE
))
#> Found more than one class "phylo" in cache; using the first, from namespace 'phyloseq'
#> Also defined by ‘tidytree’
ggtree::ggtree(tree_wo_singletons) +
ggtree::geom_nodelab(size = 2, nudge_x = -0.2, nudge_y = 0.6) +
ggtree::geom_tiplab()
#> Found more than one class "phylo" in cache; using the first, from namespace 'phyloseq'
#> Also defined by ‘tidytree’
# Without internal node singletons and only until the Genus level
tree_wo_singletons <-
phyloseq::phy_tree(taxo2tree(data_fungi_mini,
ranks = c(
"Domain", "Phylum", "Class", "Order",
"Family", "Genus"
),
internal_node_singletons = FALSE
))
#> Found more than one class "phylo" in cache; using the first, from namespace 'phyloseq'
#> Also defined by ‘tidytree’
ggtree::ggtree(tree_wo_singletons) +
ggtree::geom_nodelab(size = 2, nudge_x = -0.2, nudge_y = 0.6) +
ggtree::geom_tiplab()
#> Found more than one class "phylo" in cache; using the first, from namespace 'phyloseq'
#> Also defined by ‘tidytree’
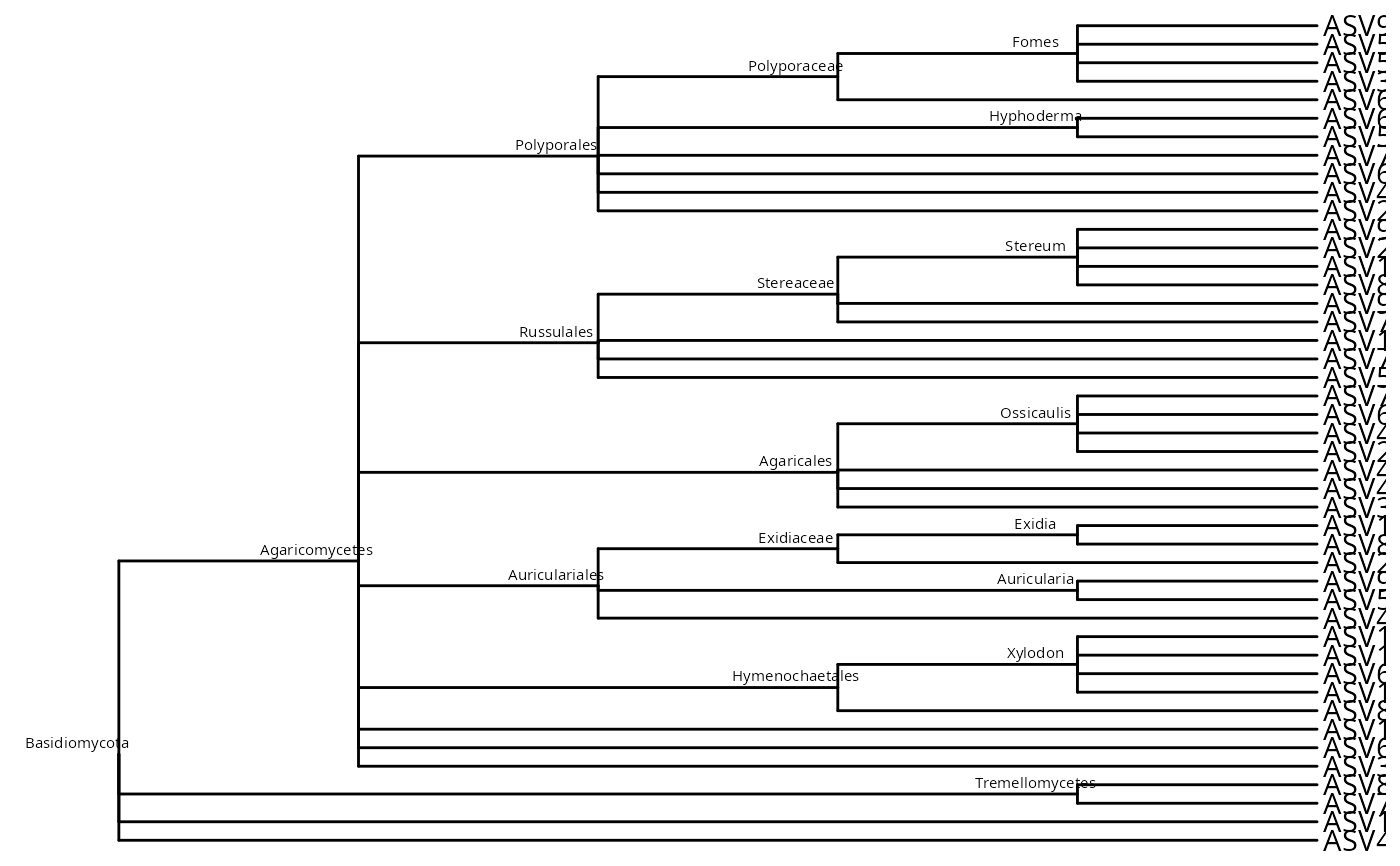 # Without taxa names (collapse identical paths)
tree_no_taxa <- taxo2tree(data_fungi_mini,
ranks = c("Domain", "Phylum", "Class", "Order", "Family", "Genus"),
use_taxa_names = FALSE
)
#> Found more than one class "phylo" in cache; using the first, from namespace 'phyloseq'
#> Also defined by ‘tidytree’
ggtree::ggtree(tree_no_taxa) +
ggtree::geom_nodelab(size = 2, nudge_x = -0.2, nudge_y = 0.6) +
ggtree::geom_tiplab()
#> Found more than one class "phylo" in cache; using the first, from namespace 'phyloseq'
#> Also defined by ‘tidytree’
# Without taxa names (collapse identical paths)
tree_no_taxa <- taxo2tree(data_fungi_mini,
ranks = c("Domain", "Phylum", "Class", "Order", "Family", "Genus"),
use_taxa_names = FALSE
)
#> Found more than one class "phylo" in cache; using the first, from namespace 'phyloseq'
#> Also defined by ‘tidytree’
ggtree::ggtree(tree_no_taxa) +
ggtree::geom_nodelab(size = 2, nudge_x = -0.2, nudge_y = 0.6) +
ggtree::geom_tiplab()
#> Found more than one class "phylo" in cache; using the first, from namespace 'phyloseq'
#> Also defined by ‘tidytree’