Estimation statistics for categorical comparisons on a list_phyloseq
Source:R/estim_lpq.R
estim_diff_lpq.RdApplies estim_diff_pq() to each phyloseq object in a list_phyloseq
and combines the results. This allows comparing estimation statistics
across different bioinformatic pipelines or parameter settings.
Arguments
- x
(required) A list_phyloseq object.
- fact
(character, required) The name of a categorical column in
sample_datato use as the grouping factor. Must be present in all phyloseq objects.- ...
Additional arguments passed to
estim_diff_pq().- verbose
(logical, default TRUE) If TRUE, print progress messages.
Value
A list of class "estim_diff_lpq_result" with components:
- results
A named list of
estim_diff_pq_resultobjects (one per phyloseq)- summary
A tibble combining all summaries with an additional
namecolumn identifying the source phyloseq
Examples
lpq <- list_phyloseq(
list(
fungi = data_fungi,
fungi_clust = postcluster_pq(data_fungi),
fungi_rarefy = rarefy_even_depth(data_fungi),
fungi_with_less_otu_in_High = multiply_counts_pq(data_fungi,
fact = "Height", prop=0.8,
conditions = "High",
multipliers = 0)
),
same_bioinfo_pipeline = FALSE
)
#> Partitioning sequences by 6-mer similarity:
#>
|
| | 0%
|
|= | 1%
|
|= | 2%
|
|== | 3%
|
|=== | 4%
|
|==== | 5%
|
|==== | 6%
|
|===== | 7%
|
|====== | 8%
|
|====== | 9%
|
|======= | 10%
|
|======== | 11%
|
|======== | 12%
|
|========= | 13%
|
|========== | 14%
|
|========== | 15%
|
|=========== | 16%
|
|============ | 17%
|
|============= | 18%
|
|============= | 19%
|
|============== | 20%
|
|=============== | 21%
|
|=============== | 22%
|
|================ | 23%
|
|================= | 24%
|
|================== | 25%
|
|================== | 26%
|
|=================== | 27%
|
|==================== | 28%
|
|==================== | 29%
|
|===================== | 30%
|
|====================== | 31%
|
|====================== | 32%
|
|======================= | 33%
|
|======================== | 34%
|
|======================== | 35%
|
|========================= | 36%
|
|========================== | 37%
|
|=========================== | 38%
|
|=========================== | 39%
|
|============================ | 40%
|
|============================= | 41%
|
|============================= | 42%
|
|============================== | 43%
|
|=============================== | 44%
|
|================================ | 45%
|
|================================ | 46%
|
|================================= | 47%
|
|================================== | 48%
|
|================================== | 49%
|
|=================================== | 50%
|
|==================================== | 51%
|
|==================================== | 52%
|
|===================================== | 53%
|
|====================================== | 54%
|
|====================================== | 55%
|
|======================================= | 56%
|
|======================================== | 57%
|
|========================================= | 58%
|
|========================================= | 59%
|
|========================================== | 60%
|
|=========================================== | 61%
|
|=========================================== | 62%
|
|============================================ | 63%
|
|============================================= | 64%
|
|============================================== | 65%
|
|============================================== | 66%
|
|=============================================== | 67%
|
|================================================ | 68%
|
|================================================ | 69%
|
|================================================= | 70%
|
|================================================== | 71%
|
|================================================== | 72%
|
|=================================================== | 73%
|
|==================================================== | 74%
|
|==================================================== | 75%
|
|===================================================== | 76%
|
|====================================================== | 77%
|
|======================================================= | 78%
|
|======================================================= | 79%
|
|======================================================== | 80%
|
|========================================================= | 81%
|
|========================================================= | 82%
|
|========================================================== | 83%
|
|=========================================================== | 84%
|
|============================================================ | 85%
|
|============================================================ | 86%
|
|============================================================= | 87%
|
|============================================================== | 88%
|
|============================================================== | 89%
|
|=============================================================== | 90%
|
|================================================================ | 91%
|
|================================================================ | 92%
|
|================================================================= | 93%
|
|================================================================== | 94%
|
|================================================================== | 95%
|
|=================================================================== | 96%
|
|==================================================================== | 97%
|
|===================================================================== | 98%
|
|===================================================================== | 99%
|
|======================================================================| 100%
#>
#> Time difference of 0.38 secs
#>
#> Sorting by relatedness within 779 groups:
#>
iteration 1 of up to 15 (100.0% stability)
iteration 2 of up to 15 (100.0% stability)
iteration 3 of up to 15 (100.0% stability)
iteration 4 of up to 15 (100.0% stability)
iteration 4 of up to 15 (100.0% stability)
#>
#> Time difference of 0.3 secs
#>
#> Clustering sequences by 9-mer similarity:
#>
|
| | 0%
|
|= | 1%
|
|= | 2%
|
|== | 3%
|
|=== | 4%
|
|==== | 5%
|
|==== | 6%
|
|===== | 7%
|
|====== | 8%
|
|====== | 9%
|
|======= | 10%
|
|======== | 11%
|
|======== | 12%
|
|========= | 13%
|
|========== | 14%
|
|========== | 15%
|
|=========== | 16%
|
|============ | 17%
|
|============= | 18%
|
|============= | 19%
|
|============== | 20%
|
|=============== | 21%
|
|=============== | 22%
|
|================ | 23%
|
|================= | 24%
|
|================== | 25%
|
|================== | 26%
|
|=================== | 27%
|
|==================== | 28%
|
|==================== | 29%
|
|===================== | 30%
|
|====================== | 31%
|
|====================== | 32%
|
|======================= | 33%
|
|======================== | 34%
|
|======================== | 35%
|
|========================= | 36%
|
|========================== | 37%
|
|=========================== | 38%
|
|=========================== | 39%
|
|============================ | 40%
|
|============================= | 41%
|
|============================= | 42%
|
|============================== | 43%
|
|=============================== | 44%
|
|================================ | 45%
|
|================================ | 46%
|
|================================= | 47%
|
|================================== | 48%
|
|================================== | 49%
|
|=================================== | 50%
|
|==================================== | 51%
|
|==================================== | 52%
|
|===================================== | 53%
|
|====================================== | 54%
|
|====================================== | 55%
|
|======================================= | 56%
|
|======================================== | 57%
|
|========================================= | 58%
|
|========================================= | 59%
|
|========================================== | 60%
|
|=========================================== | 61%
|
|=========================================== | 62%
|
|============================================ | 63%
|
|============================================= | 64%
|
|============================================== | 65%
|
|============================================== | 66%
|
|=============================================== | 67%
|
|================================================ | 68%
|
|================================================ | 69%
|
|================================================= | 70%
|
|================================================== | 71%
|
|================================================== | 72%
|
|=================================================== | 73%
|
|==================================================== | 74%
|
|==================================================== | 75%
|
|===================================================== | 76%
|
|====================================================== | 77%
|
|======================================================= | 78%
|
|======================================================= | 79%
|
|======================================================== | 80%
|
|========================================================= | 81%
|
|========================================================= | 82%
|
|========================================================== | 83%
|
|=========================================================== | 84%
|
|============================================================ | 85%
|
|============================================================ | 86%
|
|============================================================= | 87%
|
|============================================================== | 88%
|
|============================================================== | 89%
|
|=============================================================== | 90%
|
|================================================================ | 91%
|
|================================================================ | 92%
|
|================================================================= | 93%
|
|================================================================== | 94%
|
|================================================================== | 95%
|
|=================================================================== | 96%
|
|==================================================================== | 97%
|
|===================================================================== | 98%
|
|===================================================================== | 99%
|
|======================================================================| 100%
#>
#> Time difference of 4.57 secs
#>
#> Clusters via relatedness sorting: 100% (0% exclusively)
#> Clusters via rare 6-mers: 100% (0% exclusively)
#> Estimated clustering effectiveness: 100%
#>
#> You set `rngseed` to FALSE. Make sure you've set & recorded
#> the random seed of your session for reproducibility.
#> See `?set.seed`
#> ...
#> 1032OTUs were removed because they are no longer
#> present in any sample after random subsampling
#> ...
#> Modified 1136 taxa in 41 matched samples
#> ℹ Building summary table for 4 phyloseq objects...
#> ℹ Computing comparison characteristics...
#> ℹ Checking sample and taxa overlap...
#> ℹ Detected comparison type: ROBUSTNESS
#> ℹ 185 common samples, 223 common taxa
#> ✔ list_phyloseq created (ROBUSTNESS)
results <- estim_diff_lpq(lpq, fact = "Height")
#> Running estimation statistics (categorical) on 4 phyloseq objects
#> Factor: Height
#> → Processing: fungi
#> → Processing: fungi_clust
#> → Processing: fungi_rarefy
#> → Processing: fungi_with_less_otu_in_High
#> Warning: Error processing 'fungi_with_less_otu_in_High': estimated adjustment 'w' is infinite
results$summary
#> # A tibble: 18 × 9
#> name metric comparison effect_size ci_lower ci_upper pvalue_permtest
#> <chr> <chr> <chr> <dbl> <dbl> <dbl> <dbl>
#> 1 fungi Hill_0 High vs Low -0.120 -0.560 0.309 0.577
#> 2 fungi Hill_0 High vs Mi… -0.187 -0.617 0.232 0.386
#> 3 fungi Hill_1 High vs Low -0.0158 -0.434 0.418 0.942
#> 4 fungi Hill_1 High vs Mi… 0.163 -0.254 0.585 0.454
#> 5 fungi Hill_2 High vs Low -0.0387 -0.470 0.387 0.869
#> 6 fungi Hill_2 High vs Mi… 0.187 -0.230 0.592 0.396
#> 7 fungi_clust Hill_0 High vs Low -0.137 -0.580 0.289 0.520
#> 8 fungi_clust Hill_0 High vs Mi… -0.180 -0.612 0.230 0.410
#> 9 fungi_clust Hill_1 High vs Low 0.0433 -0.364 0.462 0.837
#> 10 fungi_clust Hill_1 High vs Mi… 0.221 -0.193 0.622 0.314
#> 11 fungi_clust Hill_2 High vs Low 0.0353 -0.380 0.456 0.873
#> 12 fungi_clust Hill_2 High vs Mi… 0.239 -0.171 0.640 0.276
#> 13 fungi_rarefy Hill_0 High vs Low -0.0724 -0.497 0.349 0.694
#> 14 fungi_rarefy Hill_0 High vs Mi… -0.0217 -0.434 0.413 0.877
#> 15 fungi_rarefy Hill_1 High vs Low -0.0647 -0.491 0.360 0.766
#> 16 fungi_rarefy Hill_1 High vs Mi… -0.00864 -0.429 0.426 0.971
#> 17 fungi_rarefy Hill_2 High vs Low -0.0631 -0.495 0.363 0.770
#> 18 fungi_rarefy Hill_2 High vs Mi… 0.00665 -0.418 0.435 0.976
#> # ℹ 2 more variables: pvalue_welch <dbl>, pvalue_mann_whitney <dbl>
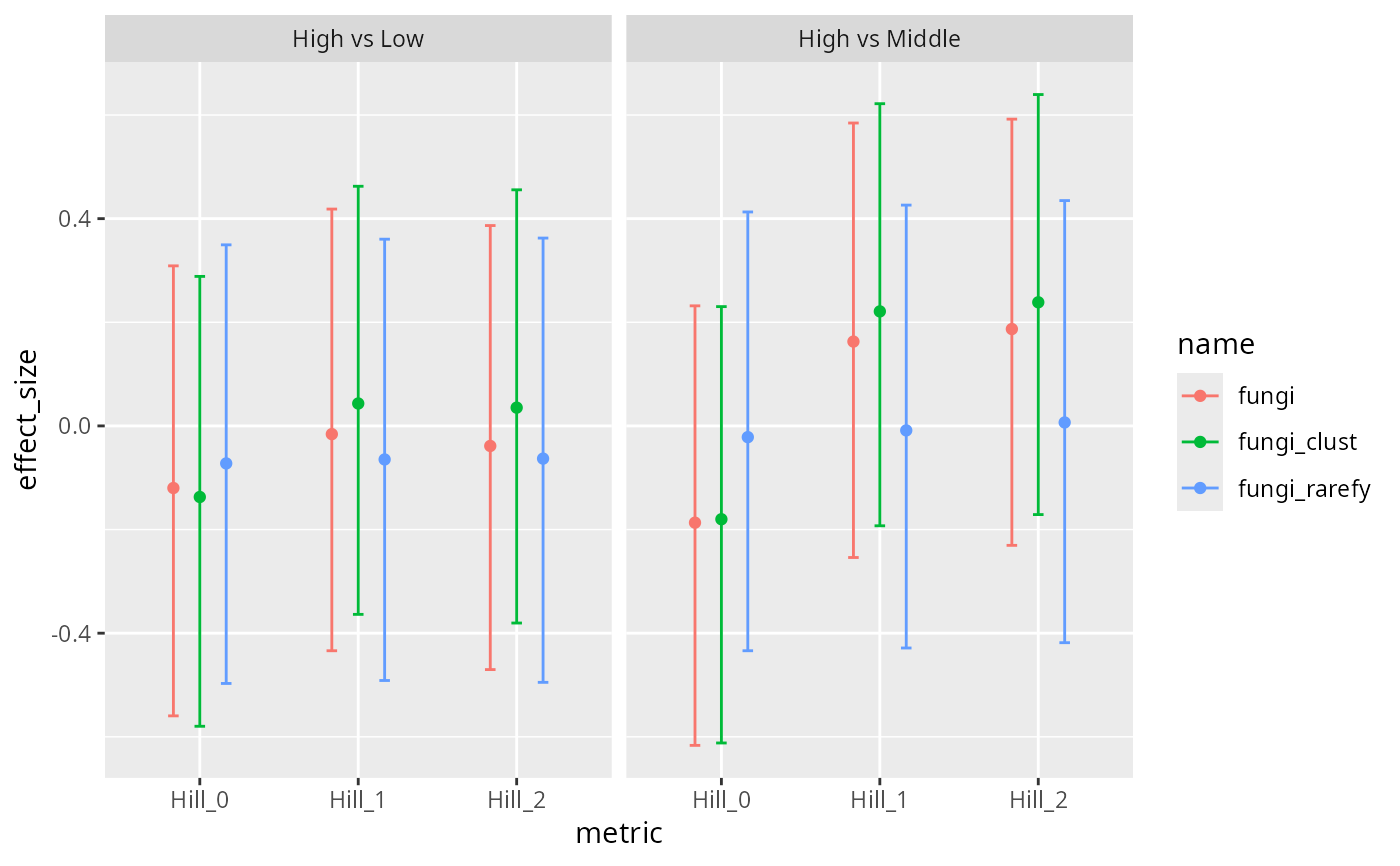
# Plot results for two phyloseq objects
ggplot(results$summary, aes(x = metric, y = effect_size, color=name)) +
facet_wrap(~comparison) +
geom_point(position=position_dodge(width=0.5)) +
geom_errorbar(aes(ymin = ci_lower, ymax = ci_upper), width = 0.2, position=position_dodge(width=0.5 ))