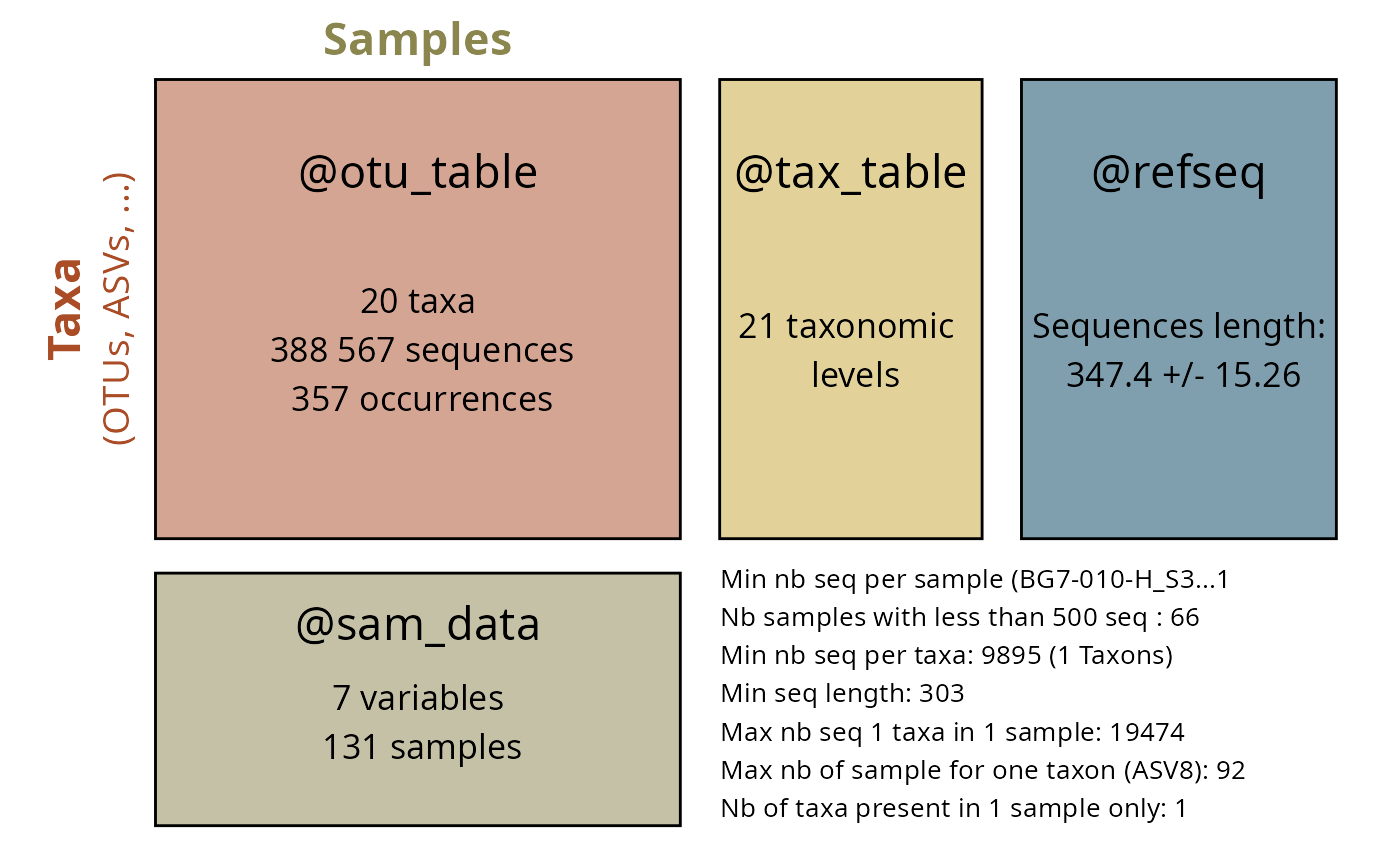
Overview
The Global Biodiversity Information Facility (GBIF) is the world’s largest repository of biodiversity occurrence data. The taxinfo package provides seamless integration with GBIF through specialized functions that retrieve occurrence data, create distribution maps, and analyze biogeographic patterns for taxa in your phyloseq objects.
Core GBIF Functions
-
tax_gbif_occur_pq(): Retrieve occurrence counts from GBIF -
plot_tax_gbif_pq(): Create distribution maps -
range_bioreg_pq(): Analyze biogeographic ranges -
tax_check_ecoregion(): Validate occurrences against ecoregions -
tax_occur_check_pq(): Check occurrences within a geographic radius
Note: Functions like
tax_gbif_occur_pq() and tax_occur_check_pq()
can work with either phyloseq objects or vectors of taxonomic names
(taxnames parameter). When using a phyloseq object, results
are automatically added to the tax_table. When using
taxnames, a tibble is returned.
Basic Occurrence Data Retrieval
Get verified taxonomic names
# Load and prepare example data
data("data_fungi_mini", package = "MiscMetabar")
# Keep only first 20 taxa for speed
data_clean <- prune_taxa(taxa = taxa_names(data_fungi_mini)[1:20], data_fungi_mini) |>
gna_verifier_pq(data_sources = 210)Find GBIF occurrence counts and add to phyloseq object
data_clean_gbif <- tax_gbif_occur_pq(data_clean)Occurrence Counts by Country
Analyze geographic distribution patterns:
data_clean_gbif_country <- tax_gbif_occur_pq(data_clean, by_country = TRUE)
data_clean_gbif_country@tax_table |>
as.data.frame() |>
select(currentCanonicalSimple, US, FR, DE, GB, ES, MX) |>
tidyr::pivot_longer(
cols = c(US, FR, DE, GB, ES, MX),
names_to = "country",
values_to = "occurrences"
) |>
mutate(occurrences = as.numeric(occurrences)) |>
filter(!is.na(occurrences), occurrences > 0) |>
ggplot(aes(x = country, y = occurrences, fill = country)) +
geom_violin() +
geom_jitter(width = 0.2, alpha = 0.5, height = 0) +
scale_y_log10() +
stat_summary(
fun.data = function(x) {
return(data.frame(y = max(x) + 0.1, label = paste("n =", length(x))))
},
geom = "text",
vjust = 0
) +
labs(
title = "GBIF Occurrences by Country",
x = "Country",
y = "Number of occurrences (log scale)"
) +
theme_idest() +
theme(legend.position = "none")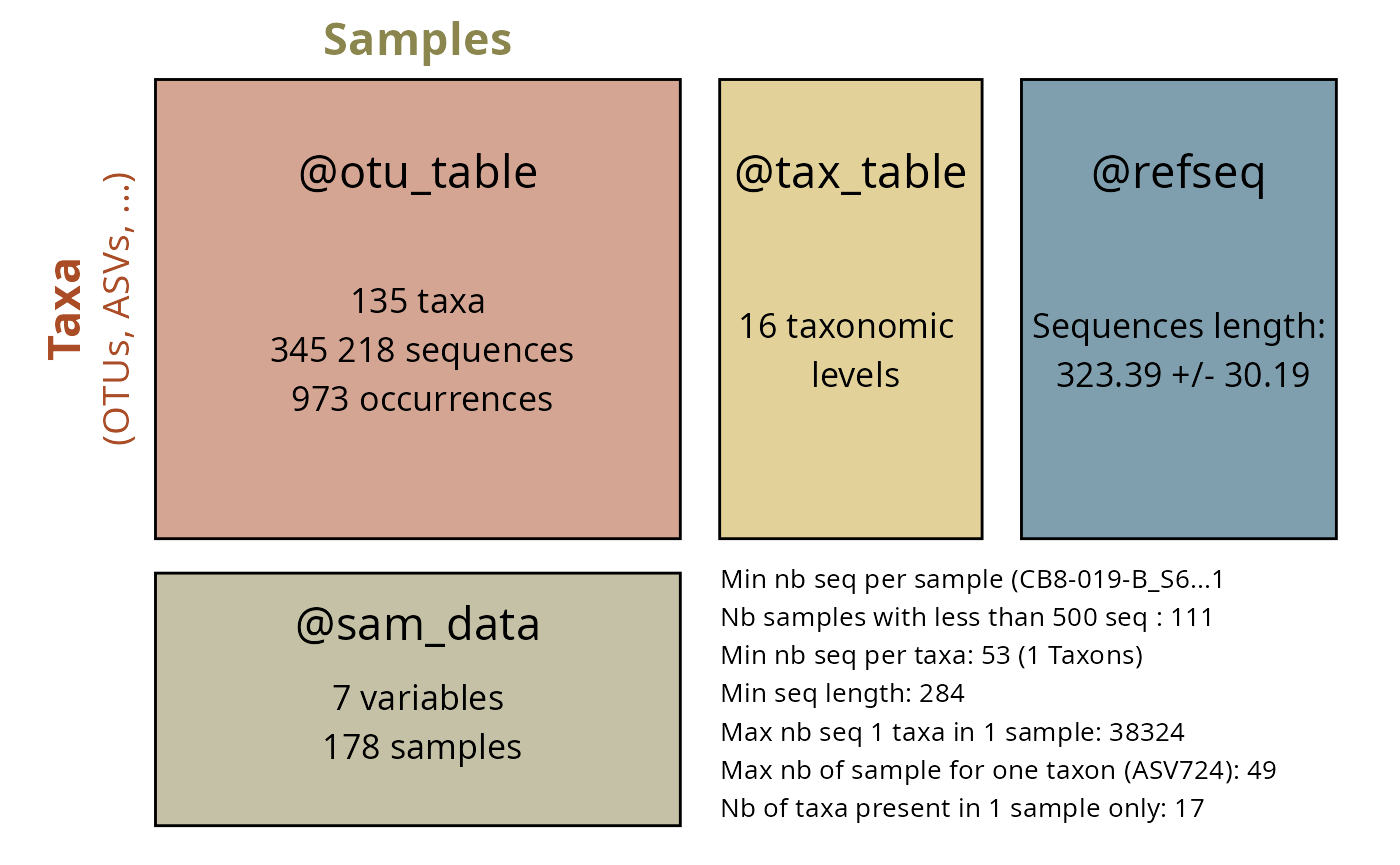
Temporal Patterns
Examine occurrence trends over time. Here we don’t use
add_to_phyloseq since the data frame output is
sufficient.
gbif_years <- tax_gbif_occur_pq(data_clean,
by_years = TRUE,
add_to_phyloseq = FALSE
) |>
tidyr::pivot_longer(cols = -canonicalName, names_to = "year", values_to = "count") |>
mutate(year = as.numeric(year))
# Visualize temporal trends for the occurences of taxa
gbif_years |>
ggplot(aes(x = year, y = log10(count + 1), color = canonicalName)) +
geom_line() +
geom_line(data = filter(gbif_years, !is.na(count)), linetype = "dotted") +
geom_point() +
labs(
title = "GBIF Occurrences Over Time",
x = "Year",
y = "Number of GBIF occurrences"
) +
theme_idest() +
ggrepel::geom_text_repel(
data = summarise(group_by(arrange(filter(gbif_years, !is.na(count)), desc(year)), canonicalName), count = first(count), year = first(year)),
aes(label = canonicalName, x = as.numeric(year)), hjust = 0,
direction = "y", size = 3
) +
xlim(2010, 2026) +
theme(legend.position = "none")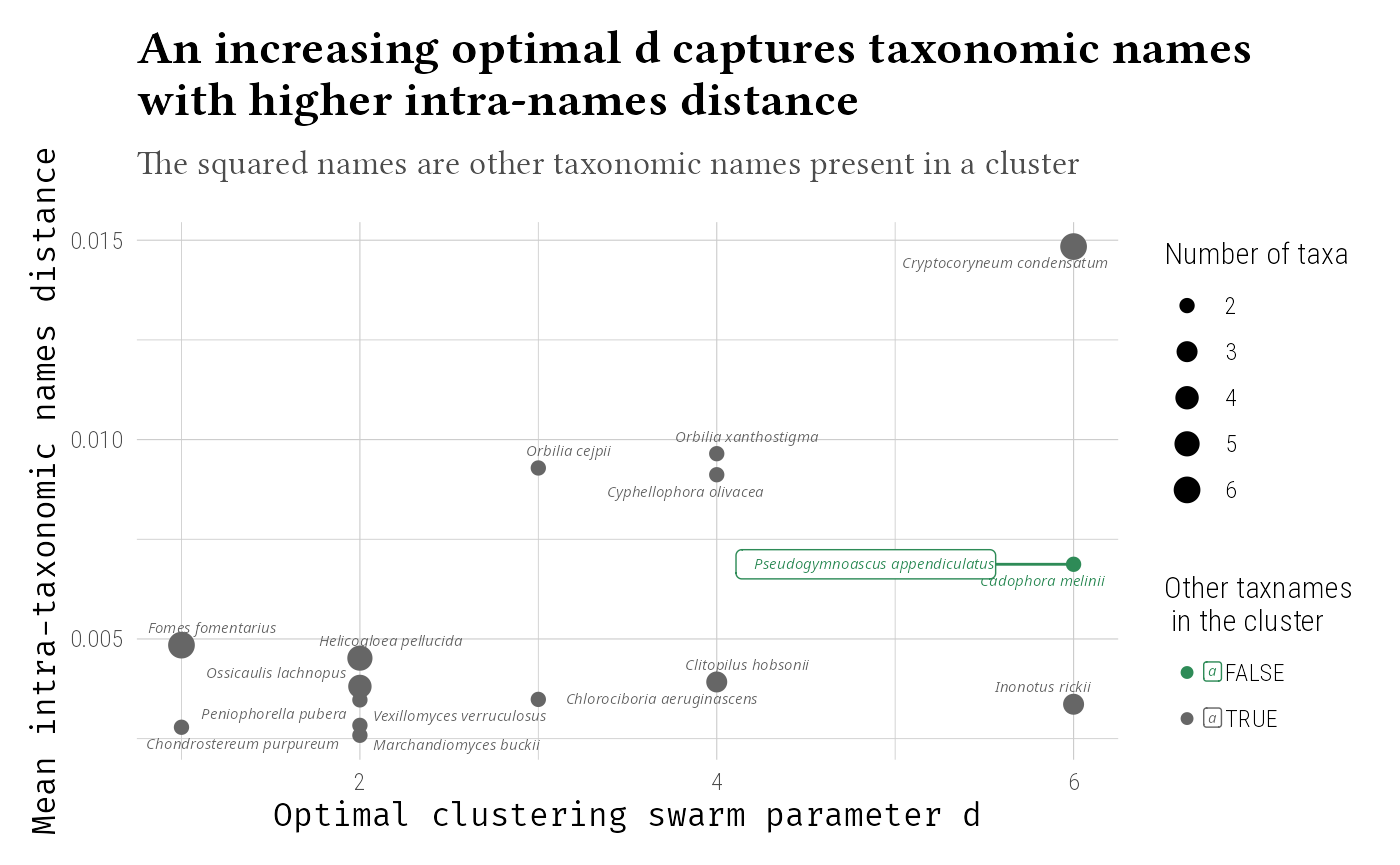
gbif_years_cumsum <- gbif_years |>
mutate(year = as.numeric(year)) |>
arrange(year) |>
group_by(canonicalName) |>
mutate(year = as.numeric(year)) |>
mutate(cum_count = cumsum(tidyr::replace_na(count, 0)))
ggplot(gbif_years_cumsum, aes(
x = year, y = log10(cum_count + 1),
color = canonicalName
)) +
geom_line() +
geom_point() +
labs(
title = "GBIF Cumulative Occurrences Over Time",
x = "Year",
y = "Number of GBIF occurrences"
) +
theme_idest() +
ggrepel::geom_text_repel(
data = summarise(
group_by(
arrange(gbif_years_cumsum, desc(year)),
canonicalName
),
cum_count = first(cum_count),
year = first(year)
),
aes(
label = canonicalName,
x = as.numeric(year)
),
hjust = 0, direction = "y", size = 3
) +
xlim(2008, 2026) +
theme(legend.position = "none")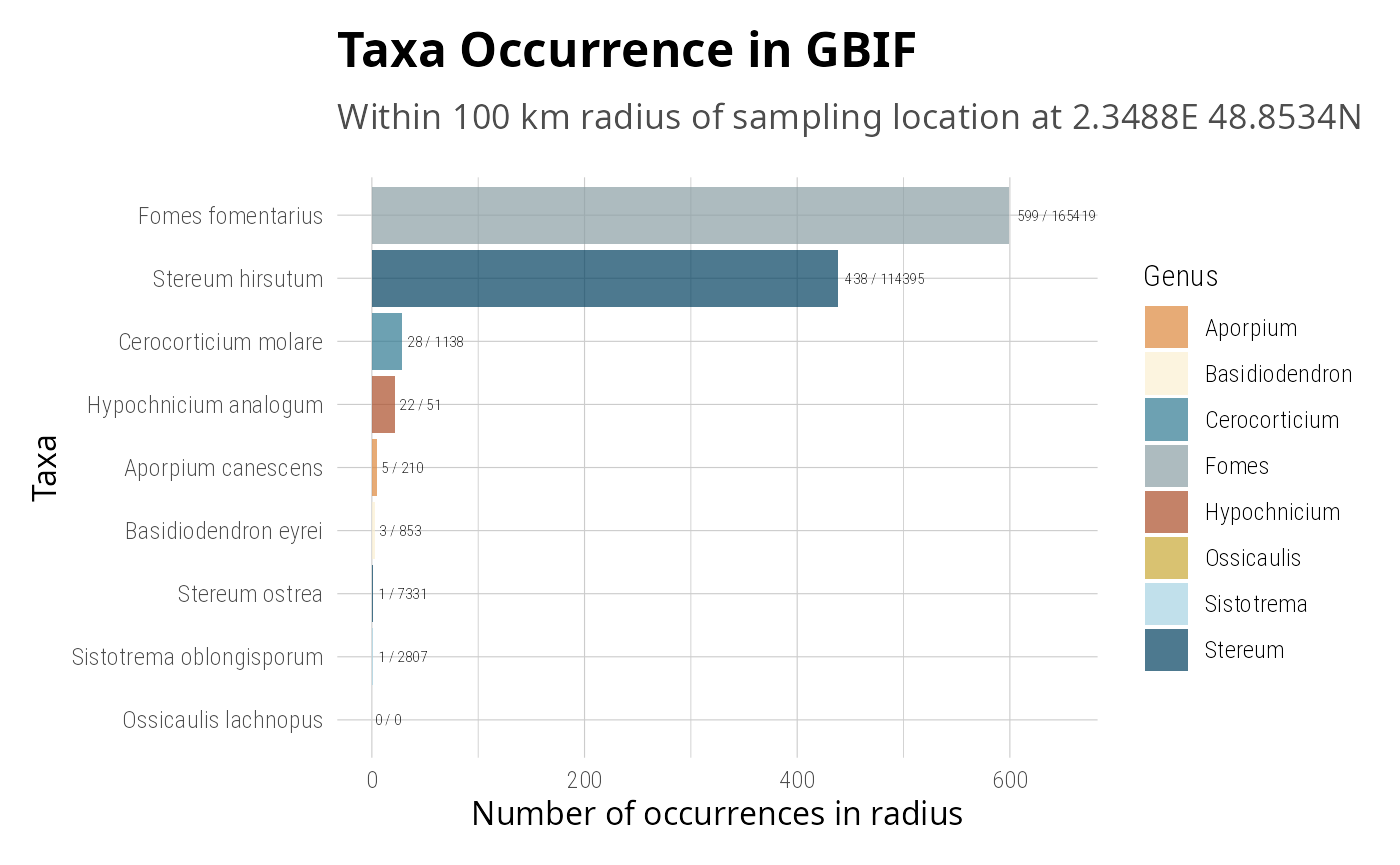
Visualization and Mapping
Basic Distribution Plots
Create occurrence distribution visualizations:
# Plot global vs regional occurrences
psmelt(data_clean_gbif) |>
as.data.frame() |>
group_by(currentCanonicalSimple) |>
summarise(
Global_occurences = as.numeric(unique(Global_occurences)),
nb_seq = sum(Abundance),
nb_samp = sum(Abundance > 0)
) |>
filter(!is.na(Global_occurences)) |>
ggplot(aes(
x = log10(1 + nb_seq),
y = log10(1 + Global_occurences)
)) +
geom_point(alpha = 0.7) +
ggrepel::geom_text_repel(aes(label = currentCanonicalSimple),
vjust = -0.5,
size = 3,
fontface = "italic",
min.segment.length = 0.2,
force = 4
) +
coord_flip() +
labs(
title = "GBIF Occurrence Counts vs molecular abundance",
x = "Number of sequences (log scale)",
y = "Number of GBIF occurrences (log scale)"
) +
theme_idest()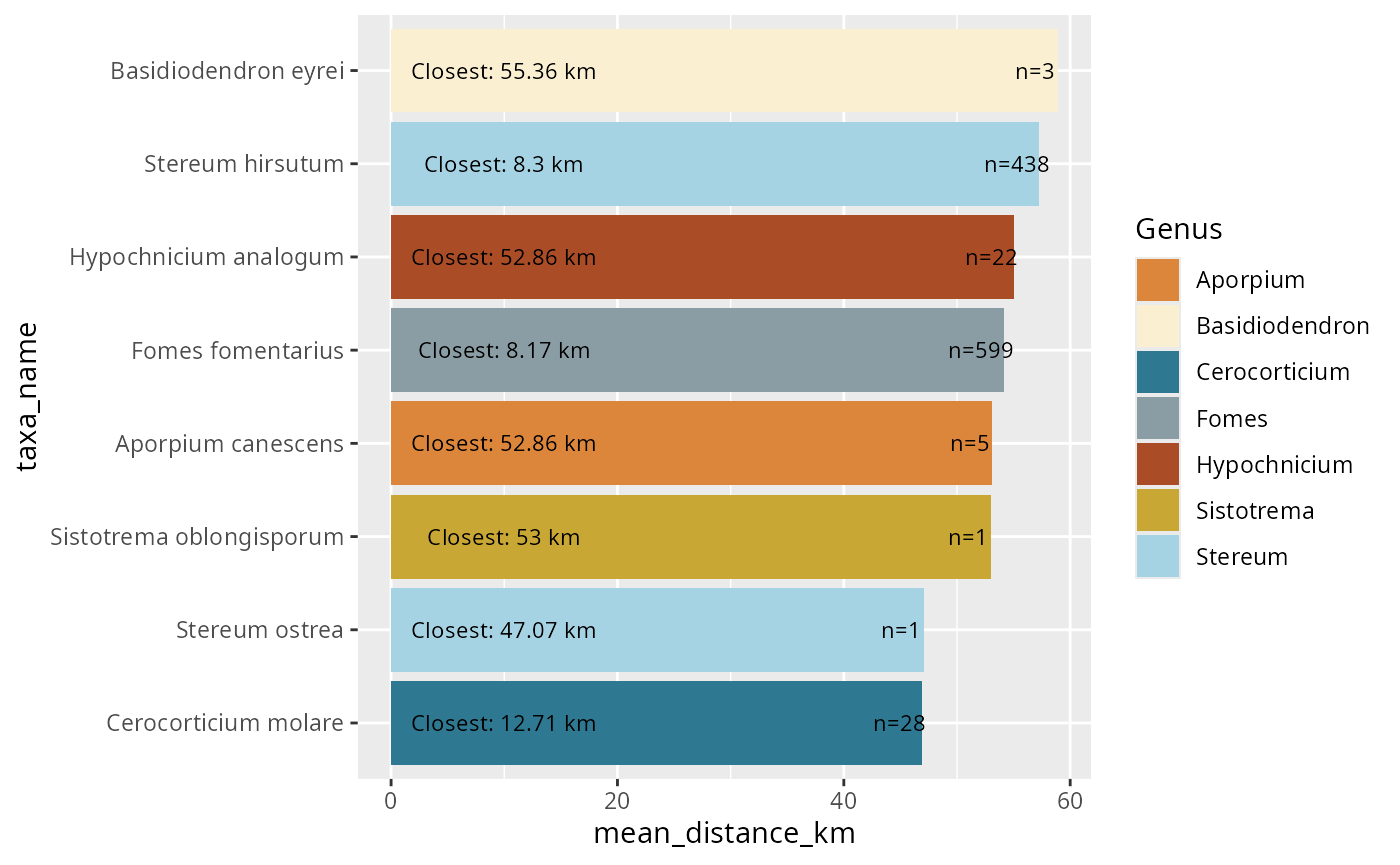
Interactive Distribution Maps
Create interactive maps showing species distributions:
plot_tax_gbif_pq(select_taxa_pq(data_clean, taxnames = "Ossicaulis lachnopus"),
interactive_plot = TRUE
)#> >>>>>>>> Total number of records: 191
#> ...GBIF records of Ossicaulis lachnopus : download of all records starting...
#> ----------------- 100 %...
#> ---> Grain filtering...
#> Records removed: 2
#> ---> Removal of duplicated records...
#> Records removed: 71
#> ---> Removal of absence records...
#> Records removed: 0
#> ---> Basis of records selection...
#> Records removed: 45
#> ---> Establishment of records selection...
#> Records removed: 0
#> ---> Time period selection...
#> Records removed: 0
#> ---> Removal of identical xy records...
#> Records removed: 0
#> ---> Removal of raster centroids...
#> Records removed: 0#> [[1]]Static Distribution Maps
distribution_maps <- plot_tax_gbif_pq(
taxnames = c("Ossicaulis lachnopus", "Basidiodendron eyrei"),
n_occur = 1000
)#> >>>>>>>> Total number of records: 191
#> ...GBIF records of Ossicaulis lachnopus : download of all records starting...
#> ----------------- 100 %...
#> ---> Grain filtering...
#> Records removed: 2
#> ---> Removal of duplicated records...
#> Records removed: 71
#> ---> Removal of absence records...
#> Records removed: 0
#> ---> Basis of records selection...
#> Records removed: 45
#> ---> Establishment of records selection...
#> Records removed: 0
#> ---> Time period selection...
#> Records removed: 0
#> ---> Removal of identical xy records...
#> Records removed: 0
#> ---> Removal of raster centroids...
#> Records removed: 0#> >>>>>>>> Total number of records: 815
#> ...GBIF records of Basidiodendron eyrei : download of all records starting...
#> ----------------- 100 %...
#> ---> Grain filtering...
#> Records removed: 3
#> ---> Removal of duplicated records...
#> Records removed: 288
#> ---> Removal of absence records...
#> Records removed: 0
#> ---> Basis of records selection...
#> Records removed: 302
#> ---> Establishment of records selection...
#> Records removed: 0
#> ---> Time period selection...
#> Records removed: 0
#> ---> Removal of identical xy records...
#> Records removed: 0
#> ---> Removal of raster centroids...
#> Records removed: 0
distribution_maps[[1]]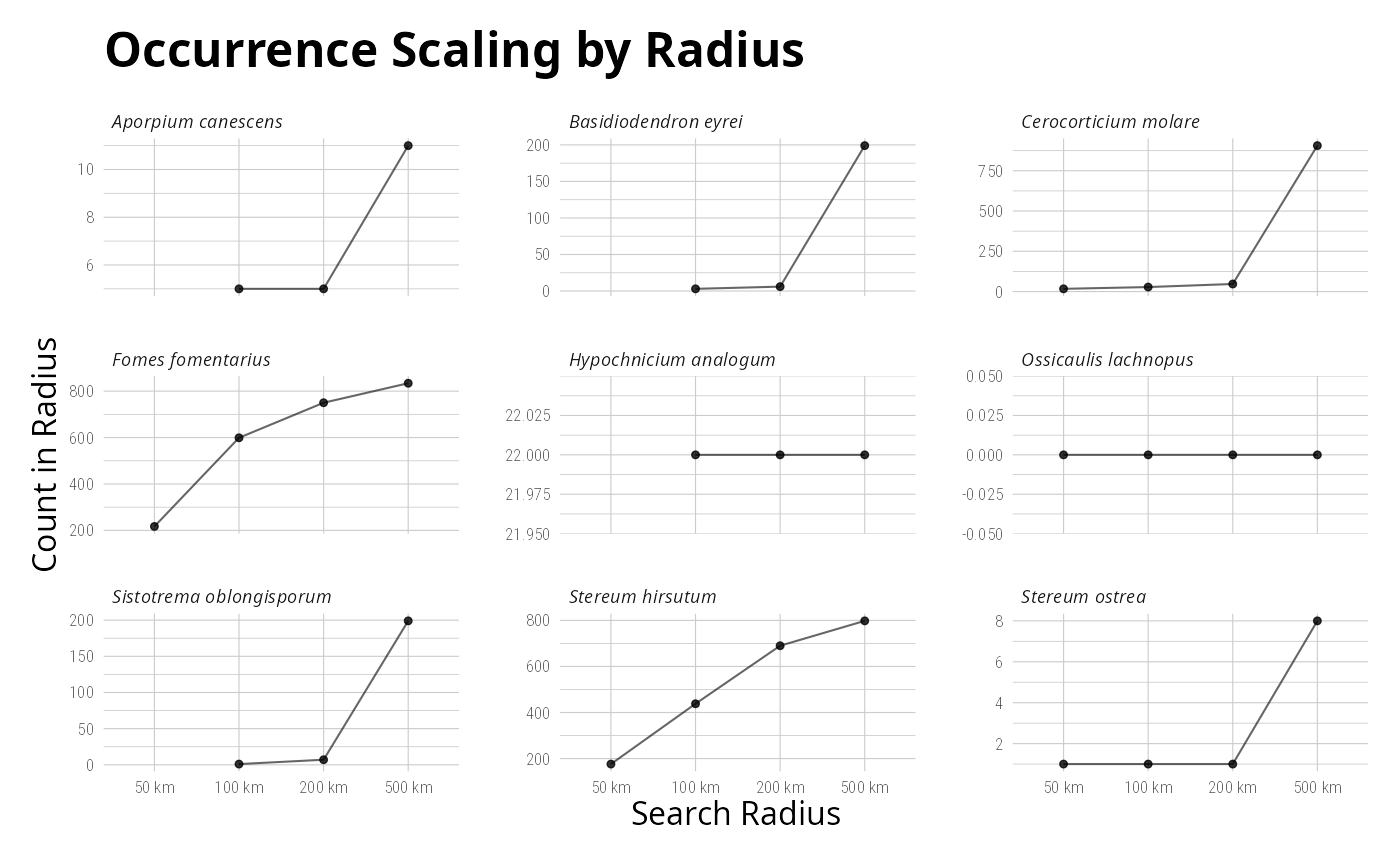
distribution_maps[[2]]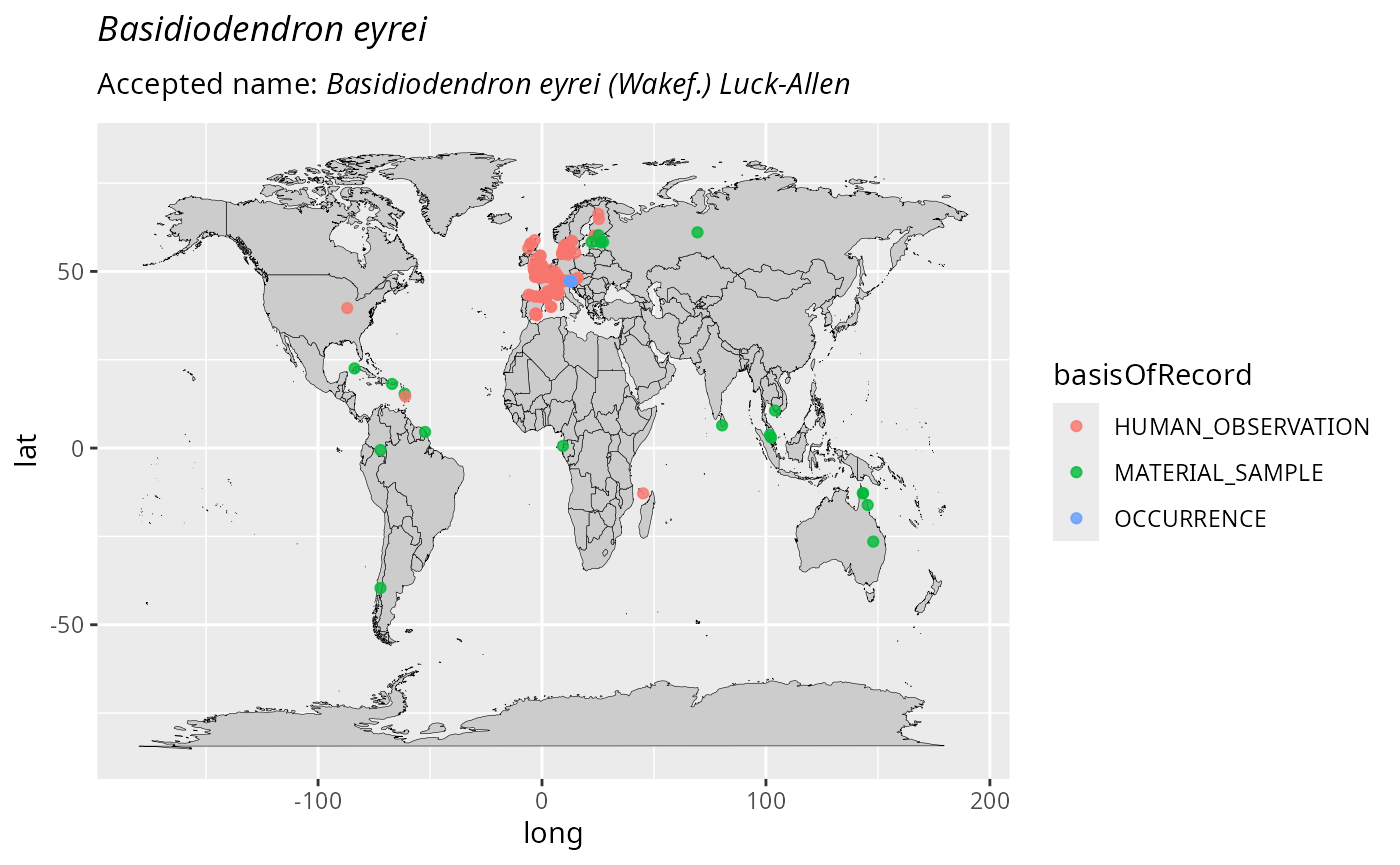
Biogeographic Analysis
Range Analysis
data_clean_range <- range_bioreg_pq(
select_taxa_pq(data_clean,
taxnames = c("Ossicaulis lachnopus", "Basidiodendron eyrei")
),
make_plot = TRUE
)#> >>>>>>>> Total number of records: 191
#> ...GBIF records of Ossicaulis lachnopus : download of sample of records starting...
#> ----------------- 100 %...
#> ---> Grain filtering...
#> Records removed: 2
#> ---> Removal of duplicated records...
#> Records removed: 71
#> ---> Removal of absence records...
#> Records removed: 0
#> ---> Basis of records selection...
#> Records removed: 45
#> ---> Establishment of records selection...
#> Records removed: 0
#> ---> Time period selection...
#> Records removed: 0
#> ---> Removal of identical xy records...
#> Records removed: 0
#> ---> Removal of raster centroids...
#> Records removed: 0#> >>>>>>>> Total number of records: 815
#> ...GBIF records of Basidiodendron eyrei : download of sample of records starting...
#> ----------------- 100 %...
#> ---> Grain filtering...
#> Records removed: 3
#> ---> Removal of duplicated records...
#> Records removed: 288
#> ---> Removal of absence records...
#> Records removed: 0
#> ---> Basis of records selection...
#> Records removed: 302
#> ---> Establishment of records selection...
#> Records removed: 0
#> ---> Time period selection...
#> Records removed: 0
#> ---> Removal of identical xy records...
#> Records removed: 0
#> ---> Removal of raster centroids...
#> Records removed: 0
data_clean_range[[2]]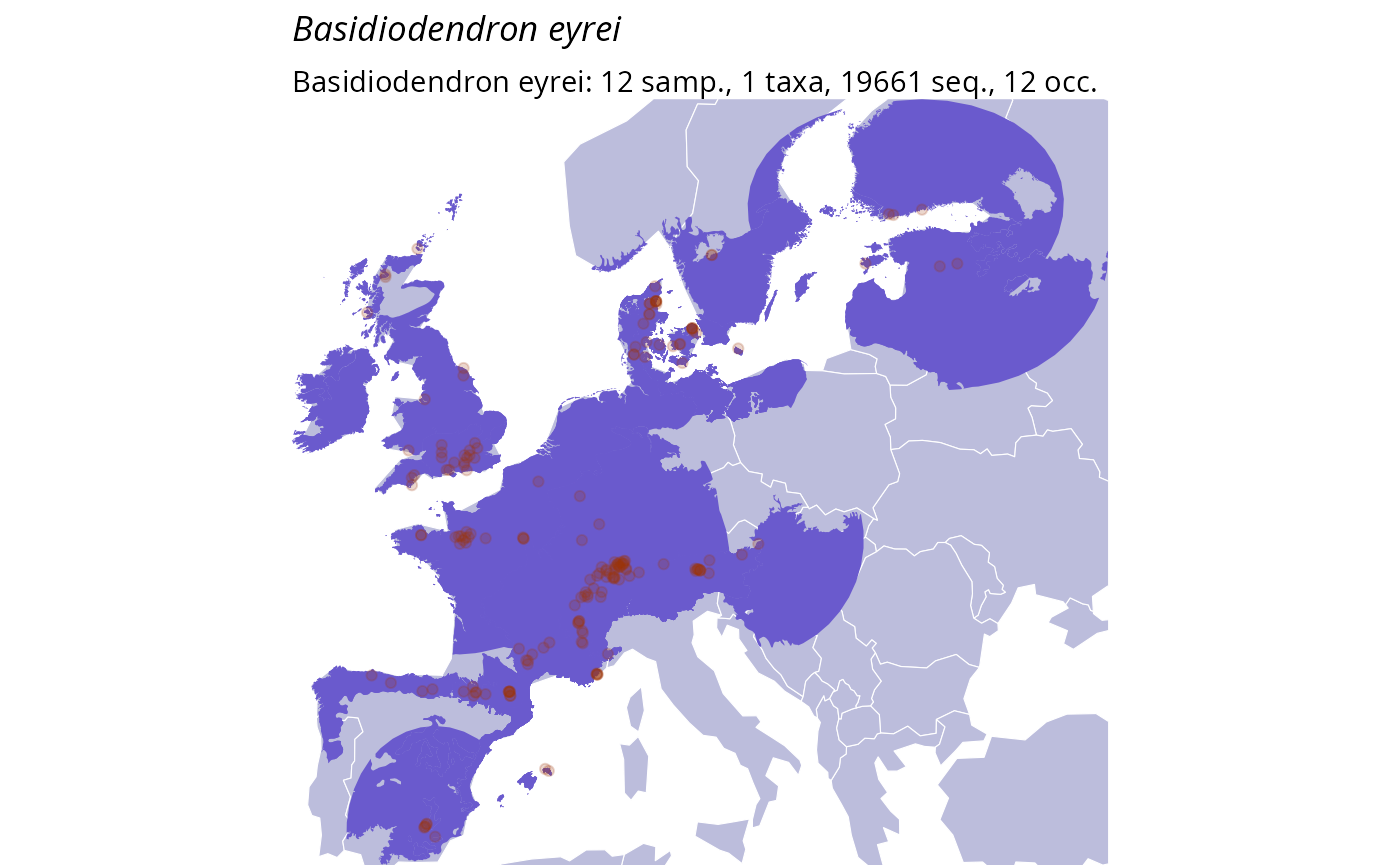
Ecoregion Validation
Check if occurrences match expected ecoregions:
tax_check_ecoregion("Xylobolus subpileatus",
longitudes = c(2.3522, 4.2),
latitudes = c(48.8566, 33)
)#> $ecoregion
#>
#> Lower New England / Northern Piedmont
#> 93
#> North Atlantic Coast
#> 44
#> Piedmont
#> 37
#> Pannonian Mixed Forests
#> 27
#> East Gulf Coastal Plain
#> 24
#> Florida Peninsula
#> 22
#> Upper East Gulf Coastal Plain
#> 22
#> Gulf Coast Prairies And Marshes
#> 18
#> Upper West Gulf Coastal Plain
#> 18
#> Cumberlands And Southern Ridge And Valley
#> 16
#> North Central Tillplain
#> 14
#> Southern Blue Ridge
#> 14
#> Great Lakes
#> 11
#> Mississippi River Alluvial Plain
#> 11
#> Ozarks
#> 11
#> South Atlantic Coastal Plain
#> 10
#> Mid-Atlantic Coastal Plain
#> 9
#> West Gulf Coastal Plain
#> 9
#> Crosstimbers And Southern Tallgrass Prairie
#> 7
#> Western Allegheny Plateau
#> 6
#> Chesapeake Bay Lowlands
#> 5
#> Interior Low Plateau
#> 5
#> Osage Plains/Flint Hills Prairie
#> 5
#> Trans-Mexican Volcanic Belt Pine-Oak Forests
#> 5
#> Central Appalachian Forest
#> 4
#> Italian Sclerophyllous And Semi-Deciduous Forests
#> 4
#> Oaxacan Montane Forests
#> 4
#> Ouachita Mountains
#> 4
#> Sierra Madre De Chiapas Moist Forests
#> 4
#> Talamancan Montane Forests
#> 4
#> Central Tallgrass Prairie
#> 3
#> Sierra Madre Oriental Pine-Oak Forests
#> 3
#> Costa Rican Seasonal Moist Forests
#> 2
#> Magdalena Valley Montane Forests
#> 2
#> Taiheiyo Evergreen Forests
#> 2
#> Alps Conifer And Mixed Forests
#> 1
#> Balsas Dry Forests
#> 1
#> Boreal Shield
#> 1
#> Cantabrian Mixed Forests
#> 1
#> Cauca Valley Montane Forests
#> 1
#> Central Mexican Matorral
#> 1
#> English Lowlands Beech Forests
#> 1
#> High Allegheny Plateau
#> 1
#> Iberian Sclerophyllous And Semi-Deciduous Forests
#> 1
#> Jalisco Dry Forests
#> 1
#> Northern Appalachian / Acadian
#> 1
#> Pindus Mountains Mixed Forests
#> 1
#> Prairie-Forest Border
#> 1
#> Sierra Madre Del Sur Pine-Oak Forests
#> 1
#> Southwest Iberian Mediterranean Sclerophyllous And Mixed Forests
#> 1
#> Superior Mixed Forest
#> 1
#>
#> $points_ecoregion
#> ECO_ID_U ECO_CODE ECO_NAME ECO_NUM
#> 1 10510 PA0402 Atlantic Mixed Forests 2
#> 2 10691 PA1321 North Saharan Steppe And Woodlands 21
#> ECODE_NAME CLS_CODE ECO_NOTES WWF_REALM
#> 1 PA0402. Atlantic mixed forests 0 <NA> PA
#> 2 PA1321. North Saharan steppe and woodlands 0 <NA> PA
#> WWF_REALM2 WWF_MHTNUM WWF_MHTNAM RealmMHT
#> 1 Palearctic 4 Temperate Broadleaf and Mixed Forests PA4
#> 2 Palearctic 13 Deserts and Xeric Shrublands PA13
#> ER_UPDATE ER_DATE_U ER_RATION SOURCEDATA
#> 1 <NA> <NA> <NA> Olson, 2001
#> 2 <NA> <NA> <NA> Olson, 2001
#>
#> $is_in_ecoregion
#> [1] FALSESession information
#> R version 4.5.1 (2025-06-13)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Kali GNU/Linux Rolling
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.29.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=fr_FR.UTF-8 LC_NUMERIC=C
#> [3] LC_TIME=fr_FR.UTF-8 LC_COLLATE=fr_FR.UTF-8
#> [5] LC_MONETARY=fr_FR.UTF-8 LC_MESSAGES=fr_FR.UTF-8
#> [7] LC_PAPER=fr_FR.UTF-8 LC_NAME=C
#> [9] LC_ADDRESS=C LC_TELEPHONE=C
#> [11] LC_MEASUREMENT=fr_FR.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: Europe/Paris
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] taxinfo_0.1.2 MiscMetabar_0.14.4 purrr_1.2.0 dplyr_1.1.4
#> [5] dada2_1.38.0 Rcpp_1.1.0 ggplot2_4.0.1 phyloseq_1.54.0
#>
#> loaded via a namespace (and not attached):
#> [1] splines_4.5.1 rnaturalearth_1.1.0
#> [3] bitops_1.0-9 urltools_1.7.3.1
#> [5] tibble_3.3.0 triebeard_0.4.1
#> [7] lifecycle_1.0.4 pwalign_1.6.0
#> [9] sf_1.0-22 lattice_0.22-7
#> [11] MASS_7.3-65 crosstalk_1.2.2
#> [13] magrittr_2.0.4 sass_0.4.10
#> [15] rmarkdown_2.30 jquerylib_0.1.4
#> [17] yaml_2.3.10 NMOF_2.11-0
#> [19] sp_2.2-0 DBI_1.2.3
#> [21] RColorBrewer_1.1-3 ade4_1.7-23
#> [23] maps_3.4.3 abind_1.4-8
#> [25] ShortRead_1.68.0 GenomicRanges_1.62.0
#> [27] BiocGenerics_0.56.0 RCurl_1.98-1.17
#> [29] satellite_1.0.6 IRanges_2.44.0
#> [31] S4Vectors_0.48.0 ggrepel_0.9.6
#> [33] gbif.range_1.0 crul_1.6.0
#> [35] terra_1.8-80 rglobi_0.3.4
#> [37] vegan_2.7-2 units_1.0-0
#> [39] brew_1.0-10 pkgdown_2.2.0
#> [41] svglite_2.2.2 permute_0.9-8
#> [43] codetools_0.2-20 DelayedArray_0.36.0
#> [45] xml2_1.5.0 tidyselect_1.2.1
#> [47] raster_3.6-32 httpcode_0.3.0
#> [49] farver_2.1.2 gmp_0.7-5
#> [51] matrixStats_1.5.0 stats4_4.5.1
#> [53] base64enc_0.1-3 Seqinfo_1.0.0
#> [55] GenomicAlignments_1.46.0 jsonlite_2.0.0
#> [57] multtest_2.66.0 e1071_1.7-16
#> [59] survival_3.8-3 iterators_1.0.14
#> [61] systemfonts_1.3.1 foreach_1.5.2
#> [63] tools_4.5.1 ragg_1.5.0
#> [65] glue_1.8.0 SparseArray_1.10.1
#> [67] leaflet.providers_2.0.0 xfun_0.54
#> [69] mgcv_1.9-4 MatrixGenerics_1.22.0
#> [71] withr_3.0.2 fastmap_1.2.0
#> [73] latticeExtra_0.6-31 rhdf5filters_1.22.0
#> [75] digest_0.6.38 CoordinateCleaner_3.0.1
#> [77] R6_2.6.1 wk_0.9.4
#> [79] textshaping_1.0.4 jpeg_0.1-11
#> [81] cigarillo_1.0.0 tidyr_1.3.1
#> [83] generics_0.1.4 data.table_1.17.8
#> [85] FNN_1.1.4.1 class_7.3-23
#> [87] httr_1.4.7 htmlwidgets_1.6.4
#> [89] S4Arrays_1.10.0 whisker_0.4.1
#> [91] pkgconfig_2.0.3 gtable_0.3.6
#> [93] S7_0.2.1 hwriter_1.3.2.1
#> [95] XVector_0.50.0 htmltools_0.5.8.1
#> [97] biomformat_1.38.0 scales_1.4.0
#> [99] tidyverse_2.0.0 Biobase_2.70.0
#> [101] ClusterR_1.3.5 png_0.1-8
#> [103] geometry_0.5.2 knitr_1.50
#> [105] rstudioapi_0.17.1 geosphere_1.5-20
#> [107] reshape2_1.4.5 rgbif_3.8.4
#> [109] uuid_1.2-1 magic_1.6-1
#> [111] nlme_3.1-168 curl_7.0.0
#> [113] proxy_0.4-27 cachem_1.1.0
#> [115] zoo_1.8-14 rhdf5_2.54.0
#> [117] stringr_1.6.0 KernSmooth_2.23-26
#> [119] parallel_4.5.1 s2_1.1.9
#> [121] desc_1.4.3 pillar_1.11.1
#> [123] grid_4.5.1 vctrs_0.6.5
#> [125] mapview_2.11.4 cluster_2.1.8.1
#> [127] evaluate_1.0.5 oai_0.4.0
#> [129] cli_3.6.5 taxize_0.10.0
#> [131] compiler_4.5.1 Rsamtools_2.26.0
#> [133] rlang_1.1.6 crayon_1.5.3
#> [135] leafpop_0.1.0 labeling_0.4.3
#> [137] mclust_6.1.2 interp_1.1-6
#> [139] classInt_0.4-11 plyr_1.8.9
#> [141] forcats_1.0.1 fs_1.6.6
#> [143] stringi_1.8.7 deldir_2.0-4
#> [145] BiocParallel_1.44.0 Biostrings_2.78.0
#> [147] lazyeval_0.2.2 leaflet_2.2.3
#> [149] Matrix_1.7-4 leafem_0.2.5
#> [151] Rhdf5lib_1.32.0 SummarizedExperiment_1.40.0
#> [153] igraph_2.2.1 RcppParallel_5.1.11-1
#> [155] bslib_0.9.0 ape_5.8-1