Check for taxa occurrences within a radius around samples using GBIF data
Source:R/tax_occur_check_pq.R
tax_occur_check_pq.RdThis function performs a species range check for taxa contained in a phyloseq object. The result can optionally be added to the phyloseq object's tax_table as new columns.
Usage
tax_occur_check_pq(
physeq = NULL,
taxnames = NULL,
taxonomic_rank = "currentCanonicalSimple",
longitude = NULL,
latitude = NULL,
radius_km = 50,
n_occur = 1000,
add_to_phyloseq = NULL,
col_prefix = NULL,
verbose = TRUE,
discard_genus_alone = taxonomic_rank == "currentCanonicalSimple",
discard_NA = TRUE,
...
)Arguments
- physeq
(optional) phyloseq object. Either `physeq` or `taxnames` must be provided, but not both. The phyloseq object containing the taxa to check.
- taxnames
(optional) A character vector of taxonomic names.
- taxonomic_rank
Character. The taxonomic rank to use for the check. Default is "currentCanonicalSimple" which corresponds to the cleaned scientific names in the phyloseq object if [gna_verifier_pq()] was used with default parameter.
- longitude
Numeric. Longitude of the test point in decimal degrees.
- latitude
Numeric. Latitude of the test point in decimal degrees.
- radius_km
Numeric. Search radius in kilometers (default: 50).
- n_occur
Numeric. Maximum number of occurrences to retrieve from GBIF for each taxon (default: 1000).
- add_to_phyloseq
(Logical, default TRUE when physeq is provided, FALSE when taxnames is provided). Whether to add the results as new columns in the phyloseq object's tax_table. If TRUE, the results will be appended to the tax_table with appropriate column names. Automatically set to TRUE when a phyloseq object is provided and FALSE when taxnames is provided. Cannot be TRUE if `taxnames` is provided.
- col_prefix
A character string to be added as a prefix to the new columns names added to the tax_table slot of the phyloseq object (default: NULL).
- verbose
(Logical, default: TRUE). Whether to print progress messages.
- ...
Additional parameters passed to [tax_occur_check()].
Value
Either a data frame (if add_to_phyloseq = FALSE) or a new phyloseq object (if add_to_phyloseq = TRUE).
Examples
data_fungi_mini_cleanNames <- gna_verifier_pq(data_fungi_mini)
#> ✔ GNA verification summary:
#> • Total taxa in phyloseq: 45
#> • Taxa submitted for verification: 37
#> • Genus-level only taxa: 2
#> • Total matches found: 25
#> • Synonyms: 2 (including 2 at genus level)
#> • Accepted names: 23 (including 21 at genus level)
check_res <- tax_occur_check_pq(data_fungi_mini_cleanNames,
longitude = 2.3,
latitude = 48,
radius_km = 100,
n_occur = 50,
add_to_phyloseq = FALSE
)
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 1 occurrences remain(s)
#> - Total original: 1
#> - Retention rate: 100%
#> ✔ Found 1 occurrences for species Stereum ostrea:
#> • Closest occurrence: 48.13 km
#> Reading ne_50m_land.zip from naturalearth...
#> Warning: Species with fewer than 7 unique records will not be tested.
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 9 occurrences remain(s)
#> - Total original: 17
#> - Retention rate: 52.9%
#> ✔ Found 9 occurrences for species Xylodon raduloides:
#> • Closest occurrence: 52.12 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 48 occurrences remain(s)
#> - Total original: 50
#> - Retention rate: 96%
#> ✔ Found 43 occurrences for species Stereum hirsutum:
#> • Closest occurrence: 11.61 km
#> ! No occurrences found for Trametopsis brasiliensis
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 3 occurrences remain(s)
#> - Total original: 3
#> - Retention rate: 100%
#> ✔ Found 3 occurrences for species Basidiodendron eyrei:
#> • Closest occurrence: 50.53 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 1 occurrences remain(s)
#> - Total original: 1
#> - Retention rate: 100%
#> ✔ Found 1 occurrences for species Sistotrema oblongisporum:
#> • Closest occurrence: 54.24 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 48 occurrences remain(s)
#> - Total original: 50
#> - Retention rate: 96%
#> ✔ Found 47 occurrences for species Fomes fomentarius:
#> • Closest occurrence: 19.21 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 1 occurrences remain(s)
#> - Total original: 1
#> - Retention rate: 100%
#> ✔ Found 1 occurrences for species Mycena renati:
#> • Closest occurrence: 54.14 km
#> ! No occurrences found for Helicogloea pellucida
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 32 occurrences remain(s)
#> - Total original: 35
#> - Retention rate: 91.4%
#> ✔ Found 31 occurrences for species Radulomyces molaris:
#> • Closest occurrence: 13.37 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 27 occurrences remain(s)
#> - Total original: 28
#> - Retention rate: 96.4%
#> ✔ Found 27 occurrences for species Elmerina caryae:
#> • Closest occurrence: 50.53 km
#> ! No occurrences found for Phanerochaete livescens
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 31 occurrences remain(s)
#> - Total original: 32
#> - Retention rate: 96.9%
#> ✔ Found 31 occurrences for species Gloeohypochnicium analogum:
#> • Closest occurrence: 50.53 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 8 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 80%
#> ✔ Found 8 occurrences for species Hyphoderma roseocremeum:
#> • Closest occurrence: 53.29 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 33 occurrences remain(s)
#> - Total original: 35
#> - Retention rate: 94.3%
#> ✔ Found 33 occurrences for species Hyphoderma setigerum:
#> • Closest occurrence: 50.53 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 37 occurrences remain(s)
#> - Total original: 50
#> - Retention rate: 74%
#> ✔ Found 31 occurrences for species Trametes versicolor:
#> • Closest occurrence: 7.4 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 1 occurrences remain(s)
#> - Total original: 1
#> - Retention rate: 100%
#> ✔ Found 1 occurrences for species Peniophora versiformis:
#> • Closest occurrence: 41.11 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 49 occurrences remain(s)
#> - Total original: 50
#> - Retention rate: 98%
#> ✔ Found 48 occurrences for species Exidia glandulosa:
#> • Closest occurrence: 15.5 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 38 occurrences remain(s)
#> - Total original: 41
#> - Retention rate: 92.7%
#> ✔ Found 38 occurrences for species Peniophorella pubera:
#> • Closest occurrence: 49.56 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 46 occurrences remain(s)
#> - Total original: 50
#> - Retention rate: 92%
#> ✔ Found 43 occurrences for species Auricularia mesenterica:
#> • Closest occurrence: 13.37 km
#> ! No occurrences found for Laetisaria buckii
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 40 occurrences remain(s)
#> - Total original: 50
#> - Retention rate: 80%
#> ✔ Found 40 occurrences for species Hericium coralloides:
#> • Closest occurrence: 46.83 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 46 occurrences remain(s)
#> - Total original: 50
#> - Retention rate: 92%
#> ✔ Found 46 occurrences for species Xylodon flaviporus:
#> • Closest occurrence: 47.96 km
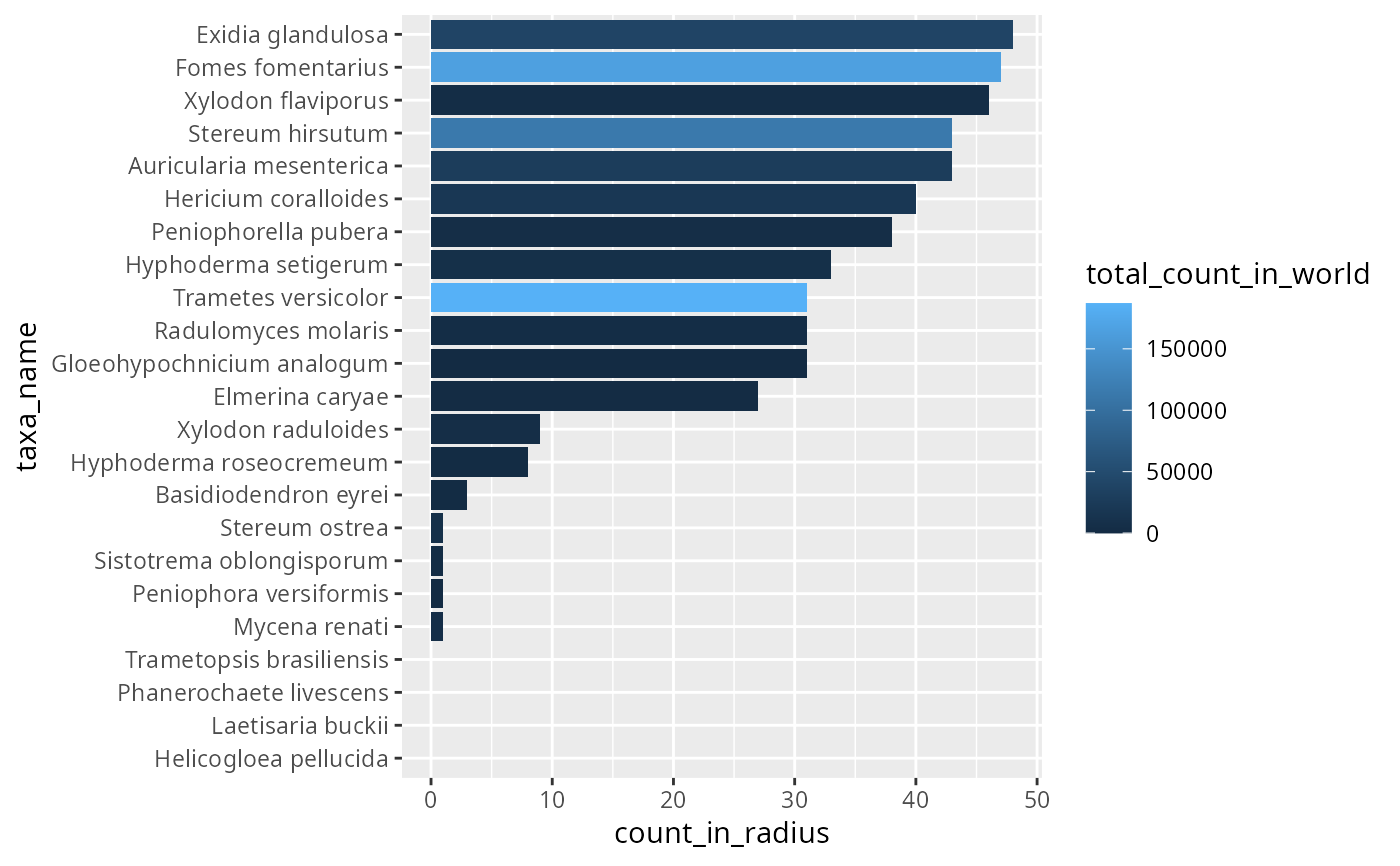
check_res |>
mutate(taxa_name = forcats::fct_reorder(taxa_name, count_in_radius)) |>
ggplot(aes(x = count_in_radius, y = taxa_name, fill = total_count_in_world)) +
geom_col()
 data_fungi_mini_cleanNames_range_verif <-
tax_occur_check_pq(data_fungi_mini_cleanNames,
longitude = 2.3,
latitude = 48,
radius_km = 50,
n_occur = 10
)
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 1 occurrences remain(s)
#> - Total original: 1
#> - Retention rate: 100%
#> ✔ Found 1 occurrences for species Stereum ostrea:
#> • Closest occurrence: 48.13 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 10 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 100%
#> ✔ Found 0 occurrences for species Xylodon raduloides:
#> • Closest occurrence: 50.53 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 10 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 100%
#> ✔ Found 7 occurrences for species Stereum hirsutum:
#> • Closest occurrence: 15.78 km
#> ! No occurrences found for Trametopsis brasiliensis
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 3 occurrences remain(s)
#> - Total original: 3
#> - Retention rate: 100%
#> ✔ Found 0 occurrences for species Basidiodendron eyrei:
#> • Closest occurrence: 50.53 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 1 occurrences remain(s)
#> - Total original: 1
#> - Retention rate: 100%
#> ✔ Found 0 occurrences for species Sistotrema oblongisporum:
#> • Closest occurrence: 54.24 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 9 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 90%
#> ✔ Found 3 occurrences for species Fomes fomentarius:
#> • Closest occurrence: 46.04 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 1 occurrences remain(s)
#> - Total original: 1
#> - Retention rate: 100%
#> ✔ Found 0 occurrences for species Mycena renati:
#> • Closest occurrence: 54.14 km
#> ! No occurrences found for Helicogloea pellucida
#> Reading ne_50m_land.zip from naturalearth...
#> Warning: Species with fewer than 7 unique records will not be tested.
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 2 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 20%
#> ✔ Found 0 occurrences for species Radulomyces molaris:
#> • Closest occurrence: 55.34 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 10 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 100%
#> ✔ Found 0 occurrences for species Elmerina caryae:
#> • Closest occurrence: 52.17 km
#> ! No occurrences found for Phanerochaete livescens
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 10 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 100%
#> ✔ Found 0 occurrences for species Gloeohypochnicium analogum:
#> • Closest occurrence: 50.53 km
#> Reading ne_50m_land.zip from naturalearth...
#> Warning: Species with fewer than 7 unique records will not be tested.
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 0 occurrences remain(s)
#> - Total original: 8
#> - Retention rate: 0%
#> ! No valid occurrences for Hyphoderma roseocremeum
#> Reading ne_50m_land.zip from naturalearth...
#> Warning: Species with fewer than 7 unique records will not be tested.
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 0 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 0%
#> ! No valid occurrences for Hyphoderma setigerum
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 10 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 100%
#> ✔ Found 4 occurrences for species Trametes versicolor:
#> • Closest occurrence: 7.4 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 1 occurrences remain(s)
#> - Total original: 1
#> - Retention rate: 100%
#> ✔ Found 1 occurrences for species Peniophora versiformis:
#> • Closest occurrence: 41.11 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 10 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 100%
#> ✔ Found 9 occurrences for species Exidia glandulosa:
#> • Closest occurrence: 20.54 km
#> Reading ne_50m_land.zip from naturalearth...
#> Warning: Species with fewer than 7 unique records will not be tested.
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 1 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 10%
#> ✔ Found 0 occurrences for species Peniophorella pubera:
#> • Closest occurrence: 54.65 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 8 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 80%
#> ✔ Found 1 occurrences for species Auricularia mesenterica:
#> • Closest occurrence: 49.8 km
#> ! No occurrences found for Laetisaria buckii
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 10 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 100%
#> ✔ Found 1 occurrences for species Hericium coralloides:
#> • Closest occurrence: 49.87 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 9 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 90%
#> ✔ Found 0 occurrences for species Xylodon flaviporus:
#> • Closest occurrence: 54.11 km
df <- data_fungi_mini_cleanNames_range_verif@tax_table[, "count_in_radius"] |>
table(useNA = "always") |>
data.frame()
colnames(df) <- c("count_in_radius", "n_taxa")
df
#> count_in_radius n_taxa
#> 1 0 6
#> 2 1 6
#> 3 3 4
#> 4 4 1
#> 5 7 1
#> 6 9 2
#> 7 <NA> 25
# Subset taxa with at least one occurrence in the radius
cond_count_sup_0 <-
data_fungi_mini_cleanNames_range_verif@tax_table[, "count_in_radius"] |>
as.numeric() > 0
cond_count_sup_0[is.na(cond_count_sup_0)] <- FALSE
names(cond_count_sup_0) <- taxa_names(data_fungi_mini_cleanNames_range_verif)
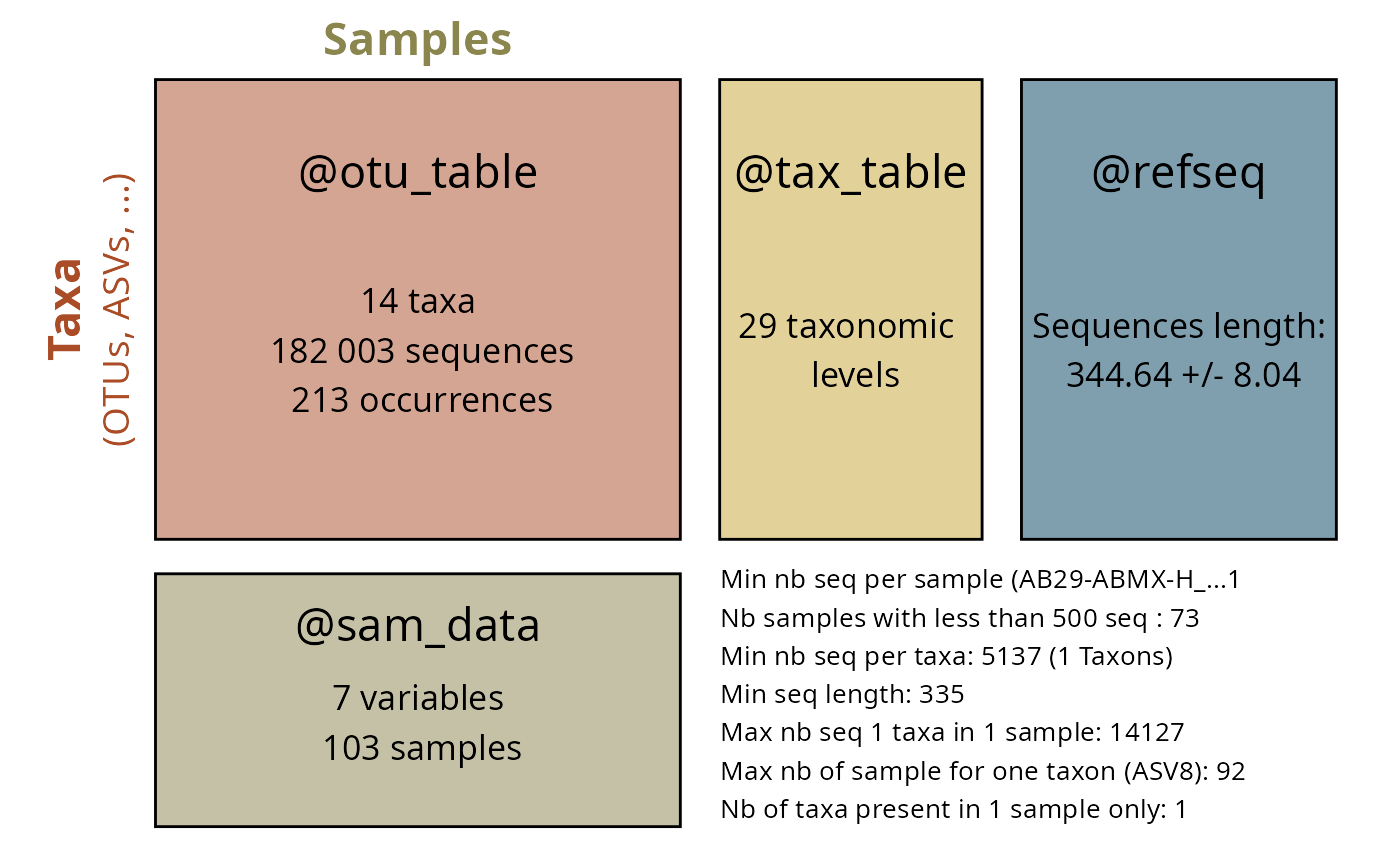
subset_taxa_pq(data_fungi_mini_cleanNames_range_verif, cond_count_sup_0) |>
summary_plot_pq()
#> Cleaning suppress 0 taxa ( ) and 34 sample(s) ( AD26-005-B_S9_MERGED.fastq.gz / AD26-005-H_S10_MERGED.fastq.gz / AD26-005-M_S11_MERGED.fastq.gz / ADABM30X-M_S16_MERGED.fastq.gz / BG7-010-H_S31_MERGED.fastq.gz / BJ8-ABM-003_S35_MERGED.fastq.gz / BQ4-018-M_S51_MERGED.fastq.gz / BR8-005_S53_MERGED.fastq.gz / BT7-006_S56_MERGED.fastq.gz / CB8-019-B_S69_MERGED.fastq.gz / CB8-019-H_S70_MERGED.fastq.gz / CB8-019-M_S71_MERGED.fastq.gz / DJ2-008-H_S88_MERGED.fastq.gz / DS1-ABM002-B_S91_MERGED.fastq.gz / DS1-ABM002-H_S92_MERGED.fastq.gz / DS1-ABM002-M_S93_MERGED.fastq.gz / DY5-004-B_S96_MERGED.fastq.gz / DY5-004-H_S97_MERGED.fastq.gz / F6-ABM-001_S105_MERGED.fastq.gz / F7-015-M_S106_MERGED.fastq.gz / H24-NVABM1-H_S111_MERGED.fastq.gz / J18-004-B_S114_MERGED.fastq.gz / J18-004-M_S116_MERGED.fastq.gz / NVABM-0163-H_S135_MERGED.fastq.gz / NVABM0216_S136_MERGED.fastq.gz / NVABM0244-M_S137_MERGED.fastq.gz / P27-ABM001_S155_MERGED.fastq.gz / T28-ABM602-B_S162_MERGED.fastq.gz / W25-ABMX_S164_MERGED.fastq.gz / W26-001-B_S165_MERGED.fastq.gz / W9-025-M_S169_MERGED.fastq.gz / X29-004-B_S174_MERGED.fastq.gz / Y28-002-B_S178_MERGED.fastq.gz / Z29-001-H_S185_MERGED.fastq.gz ).
#> Number of non-matching ASV 0
#> Number of matching ASV 45
#> Number of filtered-out ASV 31
#> Number of kept ASV 14
#> Number of kept samples 103
data_fungi_mini_cleanNames_range_verif <-
tax_occur_check_pq(data_fungi_mini_cleanNames,
longitude = 2.3,
latitude = 48,
radius_km = 50,
n_occur = 10
)
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 1 occurrences remain(s)
#> - Total original: 1
#> - Retention rate: 100%
#> ✔ Found 1 occurrences for species Stereum ostrea:
#> • Closest occurrence: 48.13 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 10 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 100%
#> ✔ Found 0 occurrences for species Xylodon raduloides:
#> • Closest occurrence: 50.53 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 10 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 100%
#> ✔ Found 7 occurrences for species Stereum hirsutum:
#> • Closest occurrence: 15.78 km
#> ! No occurrences found for Trametopsis brasiliensis
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 3 occurrences remain(s)
#> - Total original: 3
#> - Retention rate: 100%
#> ✔ Found 0 occurrences for species Basidiodendron eyrei:
#> • Closest occurrence: 50.53 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 1 occurrences remain(s)
#> - Total original: 1
#> - Retention rate: 100%
#> ✔ Found 0 occurrences for species Sistotrema oblongisporum:
#> • Closest occurrence: 54.24 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 9 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 90%
#> ✔ Found 3 occurrences for species Fomes fomentarius:
#> • Closest occurrence: 46.04 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 1 occurrences remain(s)
#> - Total original: 1
#> - Retention rate: 100%
#> ✔ Found 0 occurrences for species Mycena renati:
#> • Closest occurrence: 54.14 km
#> ! No occurrences found for Helicogloea pellucida
#> Reading ne_50m_land.zip from naturalearth...
#> Warning: Species with fewer than 7 unique records will not be tested.
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 2 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 20%
#> ✔ Found 0 occurrences for species Radulomyces molaris:
#> • Closest occurrence: 55.34 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 10 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 100%
#> ✔ Found 0 occurrences for species Elmerina caryae:
#> • Closest occurrence: 52.17 km
#> ! No occurrences found for Phanerochaete livescens
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 10 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 100%
#> ✔ Found 0 occurrences for species Gloeohypochnicium analogum:
#> • Closest occurrence: 50.53 km
#> Reading ne_50m_land.zip from naturalearth...
#> Warning: Species with fewer than 7 unique records will not be tested.
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 0 occurrences remain(s)
#> - Total original: 8
#> - Retention rate: 0%
#> ! No valid occurrences for Hyphoderma roseocremeum
#> Reading ne_50m_land.zip from naturalearth...
#> Warning: Species with fewer than 7 unique records will not be tested.
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 0 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 0%
#> ! No valid occurrences for Hyphoderma setigerum
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 10 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 100%
#> ✔ Found 4 occurrences for species Trametes versicolor:
#> • Closest occurrence: 7.4 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 1 occurrences remain(s)
#> - Total original: 1
#> - Retention rate: 100%
#> ✔ Found 1 occurrences for species Peniophora versiformis:
#> • Closest occurrence: 41.11 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 10 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 100%
#> ✔ Found 9 occurrences for species Exidia glandulosa:
#> • Closest occurrence: 20.54 km
#> Reading ne_50m_land.zip from naturalearth...
#> Warning: Species with fewer than 7 unique records will not be tested.
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 1 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 10%
#> ✔ Found 0 occurrences for species Peniophorella pubera:
#> • Closest occurrence: 54.65 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 8 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 80%
#> ✔ Found 1 occurrences for species Auricularia mesenterica:
#> • Closest occurrence: 49.8 km
#> ! No occurrences found for Laetisaria buckii
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 10 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 100%
#> ✔ Found 1 occurrences for species Hericium coralloides:
#> • Closest occurrence: 49.87 km
#> Reading ne_50m_land.zip from naturalearth...
#> ℹ After cleaning with CoordinateCleaner::clean_coordinates:
#> - 9 occurrences remain(s)
#> - Total original: 10
#> - Retention rate: 90%
#> ✔ Found 0 occurrences for species Xylodon flaviporus:
#> • Closest occurrence: 54.11 km
df <- data_fungi_mini_cleanNames_range_verif@tax_table[, "count_in_radius"] |>
table(useNA = "always") |>
data.frame()
colnames(df) <- c("count_in_radius", "n_taxa")
df
#> count_in_radius n_taxa
#> 1 0 6
#> 2 1 6
#> 3 3 4
#> 4 4 1
#> 5 7 1
#> 6 9 2
#> 7 <NA> 25
# Subset taxa with at least one occurrence in the radius
cond_count_sup_0 <-
data_fungi_mini_cleanNames_range_verif@tax_table[, "count_in_radius"] |>
as.numeric() > 0
cond_count_sup_0[is.na(cond_count_sup_0)] <- FALSE
names(cond_count_sup_0) <- taxa_names(data_fungi_mini_cleanNames_range_verif)
subset_taxa_pq(data_fungi_mini_cleanNames_range_verif, cond_count_sup_0) |>
summary_plot_pq()
#> Cleaning suppress 0 taxa ( ) and 34 sample(s) ( AD26-005-B_S9_MERGED.fastq.gz / AD26-005-H_S10_MERGED.fastq.gz / AD26-005-M_S11_MERGED.fastq.gz / ADABM30X-M_S16_MERGED.fastq.gz / BG7-010-H_S31_MERGED.fastq.gz / BJ8-ABM-003_S35_MERGED.fastq.gz / BQ4-018-M_S51_MERGED.fastq.gz / BR8-005_S53_MERGED.fastq.gz / BT7-006_S56_MERGED.fastq.gz / CB8-019-B_S69_MERGED.fastq.gz / CB8-019-H_S70_MERGED.fastq.gz / CB8-019-M_S71_MERGED.fastq.gz / DJ2-008-H_S88_MERGED.fastq.gz / DS1-ABM002-B_S91_MERGED.fastq.gz / DS1-ABM002-H_S92_MERGED.fastq.gz / DS1-ABM002-M_S93_MERGED.fastq.gz / DY5-004-B_S96_MERGED.fastq.gz / DY5-004-H_S97_MERGED.fastq.gz / F6-ABM-001_S105_MERGED.fastq.gz / F7-015-M_S106_MERGED.fastq.gz / H24-NVABM1-H_S111_MERGED.fastq.gz / J18-004-B_S114_MERGED.fastq.gz / J18-004-M_S116_MERGED.fastq.gz / NVABM-0163-H_S135_MERGED.fastq.gz / NVABM0216_S136_MERGED.fastq.gz / NVABM0244-M_S137_MERGED.fastq.gz / P27-ABM001_S155_MERGED.fastq.gz / T28-ABM602-B_S162_MERGED.fastq.gz / W25-ABMX_S164_MERGED.fastq.gz / W26-001-B_S165_MERGED.fastq.gz / W9-025-M_S169_MERGED.fastq.gz / X29-004-B_S174_MERGED.fastq.gz / Y28-002-B_S178_MERGED.fastq.gz / Z29-001-H_S185_MERGED.fastq.gz ).
#> Number of non-matching ASV 0
#> Number of matching ASV 45
#> Number of filtered-out ASV 31
#> Number of kept ASV 14
#> Number of kept samples 103
