Add FungalTraits and FUNGuild information to a phyloseq object
Source:R/fungal_traits_guilds.R
fungal_traits_guilds.RdA convenience wrapper that adds guild and trait information from both the FungalTraits database and the FUNGuild database to the `tax_table` slot of a phyloseq object. Optionally creates consensus columns that summarise agreement between the two databases.
If `currentCanonicalSimple` is not already present in the `tax_table`, [gna_verifier_pq()] is called internally to clean and verify the taxonomic names before querying the databases.
Usage
fungal_traits_guilds(
physeq,
fungal_traits_file = system.file("extdata", "fungal_traits.csv", package = "taxinfo"),
ft_taxonomic_rank = "genusEpithet",
ft_csv_rank = "GENUS",
ft_sep = "\t",
ft_col_prefix = "ft_",
fg_tax_levels = c("Kingdom", "Phylum", "Class", "Order", "Family", "Genus", "Species"),
fg_col_prefix = "fg_",
ft_csv_cols_select = c("GENUS", "COMMENT.on.genus", "primary_lifestyle",
"Secondary_lifestyle", "Comment_on_lifestyle_template",
"Endophytic_interaction_capability_template", "Plant_pathogenic_capacity_template",
"Decay_substrate_template", "Decay_type_template", "Aquatic_habitat_template",
"Animal_biotrophic_capacity_template", "Specific_hosts", "Growth_form_template",
"Fruitbody_type_template", "Hymenium_type_template",
"Ectomycorrhiza_exploration_type_template", "Ectomycorrhiza_lineage_template",
"primary_photobiont",
"secondary_photobiont"),
db_url = "http://www.stbates.org/funguild_db_2.php",
add_consensus = TRUE,
consensus_col_prefix = "cons_",
add_to_phyloseq = TRUE,
gna_data_sources = c(1, 12),
verbose = TRUE
)Arguments
- physeq
A phyloseq object.
- fungal_traits_file
(Character) Path to the FungalTraits CSV file. Defaults to the simplified version bundled with the package.
- ft_taxonomic_rank
(Character, default `"genusEpithet"`) Column in `tax_table` used to match against the FungalTraits genus column.
- ft_csv_rank
(Character, default `"GENUS"`) Column in the FungalTraits CSV file that contains genus names.
- ft_sep
(Character, default `";"`) Field separator of the FungalTraits CSV file. See [utils::read.csv()].
- ft_col_prefix
(Character, default `"ft_"`) Prefix applied to all columns imported from FungalTraits.
- fg_tax_levels
(Character vector) Names of the tax_table columns that represent the 7 standard taxonomic ranks fed to FUNGuild.
- fg_col_prefix
(Character, default `"fg_"`) Prefix applied to all columns imported from FUNGuild.
- db_url
(Character) URL of the FUNGuild database. See [MiscMetabar::get_funguild_db()].
- add_consensus
(Logical, default `TRUE`) If `TRUE`, add consensus columns comparing trophic modes assigned by the two databases.
- consensus_col_prefix
(Character, default `"cons_"`) Prefix applied to consensus columns.
- add_to_phyloseq
(Logical, default `TRUE`) If `TRUE`, return an updated phyloseq object. If `FALSE`, return a tibble of the tax_table.
- gna_data_sources
Integer or character vector passed to [gna_verifier_pq()] when taxonomic names need to be verified. See [taxize::gna_verifier()].
- verbose
(Logical, default `TRUE`) If `TRUE`, print progress messages.
Value
Either an updated phyloseq object (when `add_to_phyloseq = TRUE`) or a tibble of the augmented tax_table.
See also
[tax_info_pq()], [gna_verifier_pq()], [MiscMetabar::add_funguild_info()], [MiscMetabar::funguild_assign()]
Examples
# physeq object with already-verified names
res_guild <- data_fungi |>
gna_verifier_pq(data_sources = 210) |>
fungal_traits_guilds()
#> ✔ GNA verification summary:
#> • Total taxa in phyloseq: 1420
#> • Taxa submitted for verification: 1010
#> • Genus-level only taxa: 359
#> • Total matches found: 300
#> • Synonyms: 16 (including 15 at genus level)
#> • Accepted names: 284 (including 136 at genus level)
#> ✔ Added 17 columns from /tmp/RtmprXGs9y/temp_libpath237d77f65efda/taxinfo/extdata/fungal_traits.csv with information for 862 taxa in the tax_table slot of the phyloseq object
#> Warning: Some `fg_tax_levels` not found in tax_table: "Kingdom".
#> ✔ Added 11 FUNGuild columns to tax_table.
#> Warning: argument 'pattern' has length > 1 and only the first element will be used
#> Warning: argument 'pattern' has length > 1 and only the first element will be used
#> ✔ Added 2 consensus columns:
#> "cons_trophicMode" and "cons_trophicMode_agreement".
table(res_guild@tax_table[, "cons_trophicMode"], useNA = "always")
#>
#> Conflicting Pathotroph
#> 113 127
#> Pathotroph-Saprotroph Pathotroph-Saprotroph-Symbiotroph
#> 60 29
#> Pathotroph-Symbiotroph Saprotroph
#> 4 630
#> Saprotroph-Symbiotroph Symbiotroph
#> 31 108
#> <NA>
#> 318
table(res_guild@tax_table[, "cons_trophicMode_agreement"], useNA = "always")
#>
#> Agree Disagree Only one source <NA>
#> 745 113 562 0
# physeq object WITHOUT verified names: gna_verifier_pq is called internally
res_guild_2 <- fungal_traits_guilds(data_fungi, gna_data_sources = 210)
#> ℹ Column "currentCanonicalSimple" not found in tax_table.
#> ✔ GNA verification summary:
#> • Total taxa in phyloseq: 1420
#> • Taxa submitted for verification: 1010
#> • Genus-level only taxa: 359
#> • Total matches found: 300
#> • Synonyms: 16 (including 15 at genus level)
#> • Accepted names: 284 (including 136 at genus level)
#> ✔ Added 17 columns from /tmp/RtmprXGs9y/temp_libpath237d77f65efda/taxinfo/extdata/fungal_traits.csv with information for 862 taxa in the tax_table slot of the phyloseq object
#> Warning: Some `fg_tax_levels` not found in tax_table: "Kingdom".
#> ✔ Added 11 FUNGuild columns to tax_table.
#> Warning: argument 'pattern' has length > 1 and only the first element will be used
#> Warning: argument 'pattern' has length > 1 and only the first element will be used
#> ✔ Added 2 consensus columns:
#> "cons_trophicMode" and "cons_trophicMode_agreement".
table(res_guild_2@tax_table[, "ft_primary_lifestyle"])
#>
#> animal_endosymbiont animal_parasite
#> 4 8 25
#> dung_saprotroph ectomycorrhizal lichen_parasite
#> 3 1 14
#> lichenized litter_saprotroph mycoparasite
#> 95 92 40
#> nectar/tap_saprotroph plant_pathogen root_endophyte
#> 8 65 1
#> soil_saprotroph sooty_mold unspecified_saprotroph
#> 26 1 127
#> wood_saprotroph
#> 352
table(res_guild_2@tax_table[, "fg_trophicMode"])
#>
#> Saprotroph Symbiotroph-Saprotroph-Symbiotroph
#> 4 7
#> Pathotroph Pathotroph-Saprotroph
#> 22 256
#> Pathotroph-Saprotroph-Symbiotroph Pathotroph-Symbiotroph
#> 156 54
#> Saprotroph Saprotroph-Symbiotroph
#> 417 95
#> Symbiotroph
#> 91
table(res_guild_2@tax_table[, "cons_trophicMode"])
#>
#> Conflicting Pathotroph
#> 113 127
#> Pathotroph-Saprotroph Pathotroph-Saprotroph-Symbiotroph
#> 60 29
#> Pathotroph-Symbiotroph Saprotroph
#> 4 630
#> Saprotroph-Symbiotroph Symbiotroph
#> 31 108
# Return a tibble instead of a phyloseq
tib <- fungal_traits_guilds(data_fungi_cleanNames, add_to_phyloseq = FALSE)
#> Error: object 'data_fungi_cleanNames' not found
# \donttest{
res_guild_2 |> psmelt() |>
filter(Abundance > 0) |>
ggplot(aes(x = Height, y = Abundance, fill = cons_trophicMode)) +
geom_col() +
theme_bw() +
labs(x = "Height", y = "Molecular abundance", fill = "Consensus trophic mode") +
theme(axis.text.x = element_text(angle = 45, hjust = 1))
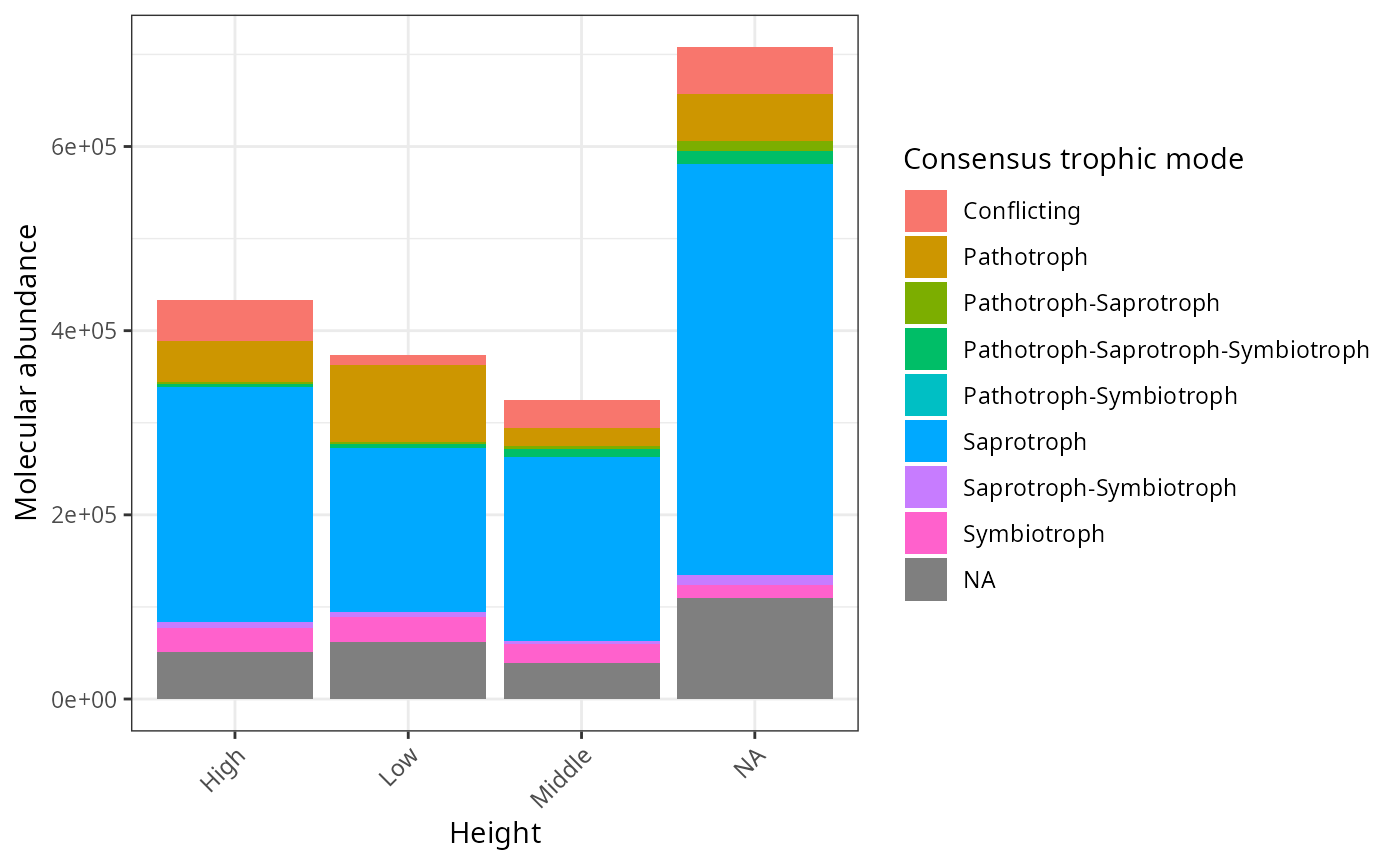 tax_bar_pq(res_guild_2,"Height", "cons_trophicMode", add_ribbon=TRUE)
tax_bar_pq(res_guild_2,"Height", "cons_trophicMode", add_ribbon=TRUE)
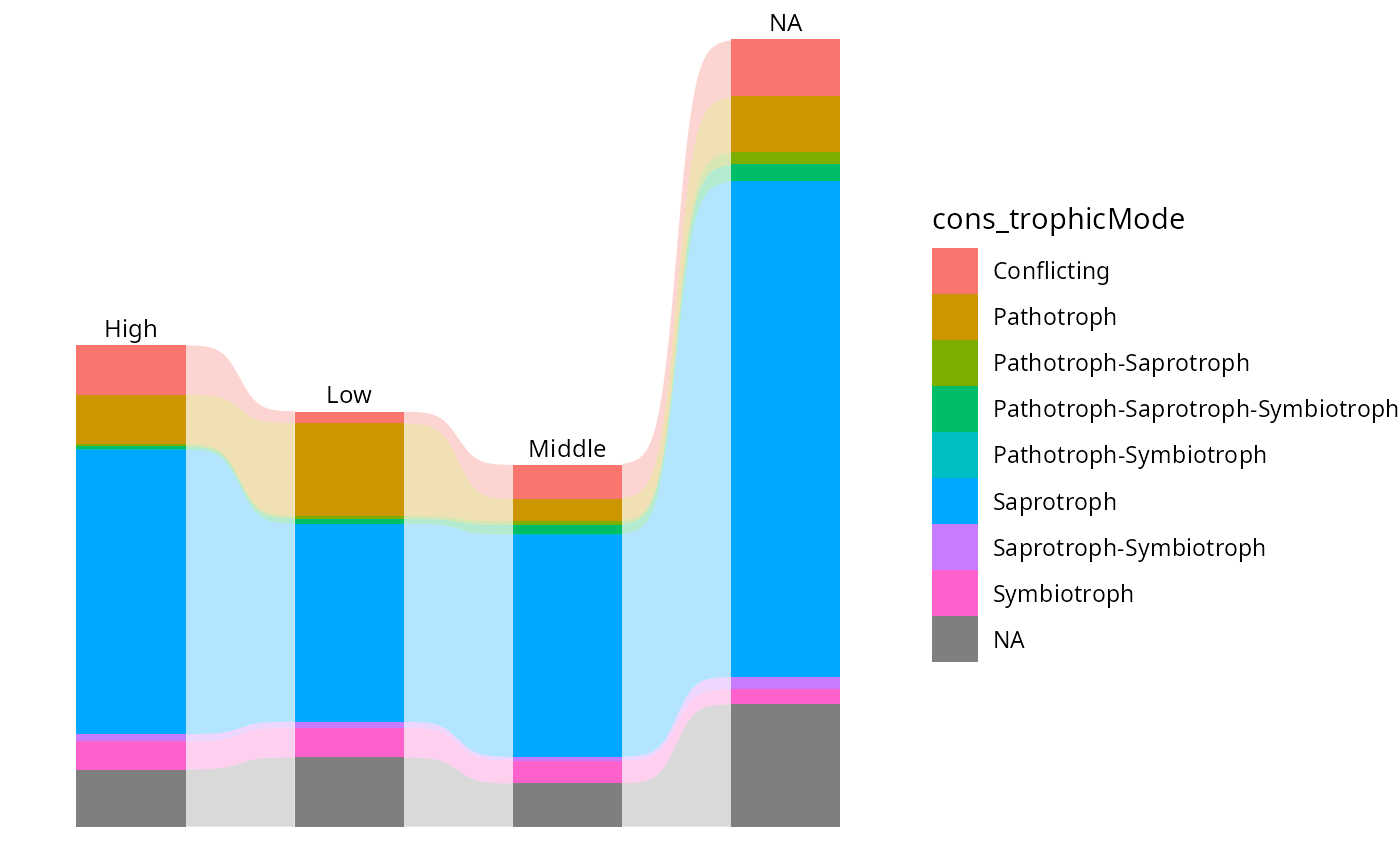 # }
# }