Add information to sample_data slot of a phyloseq-class object
Source:R/dada_phyloseq.R
add_info_to_sam_data.RdWarning: The value nb_seq and nb_otu may be outdated if you transform your
phyloseq object, e.g. using the subset_taxa_pq() function
Arguments
- physeq
(required) a
phyloseq-classobject obtained using thephyloseqpackage.- df_info
: A dataframe with rownames matching for sample names of the phyloseq object
- add_nb_seq
(Logical, default TRUE) Does we add a column nb_seq collecting the number of sequences per sample?
- add_nb_otu
(Logical, default TRUE) Does we add a column nb_otu collecting the number of OTUs per sample?
Examples
data_fungi <- add_info_to_sam_data(data_fungi)
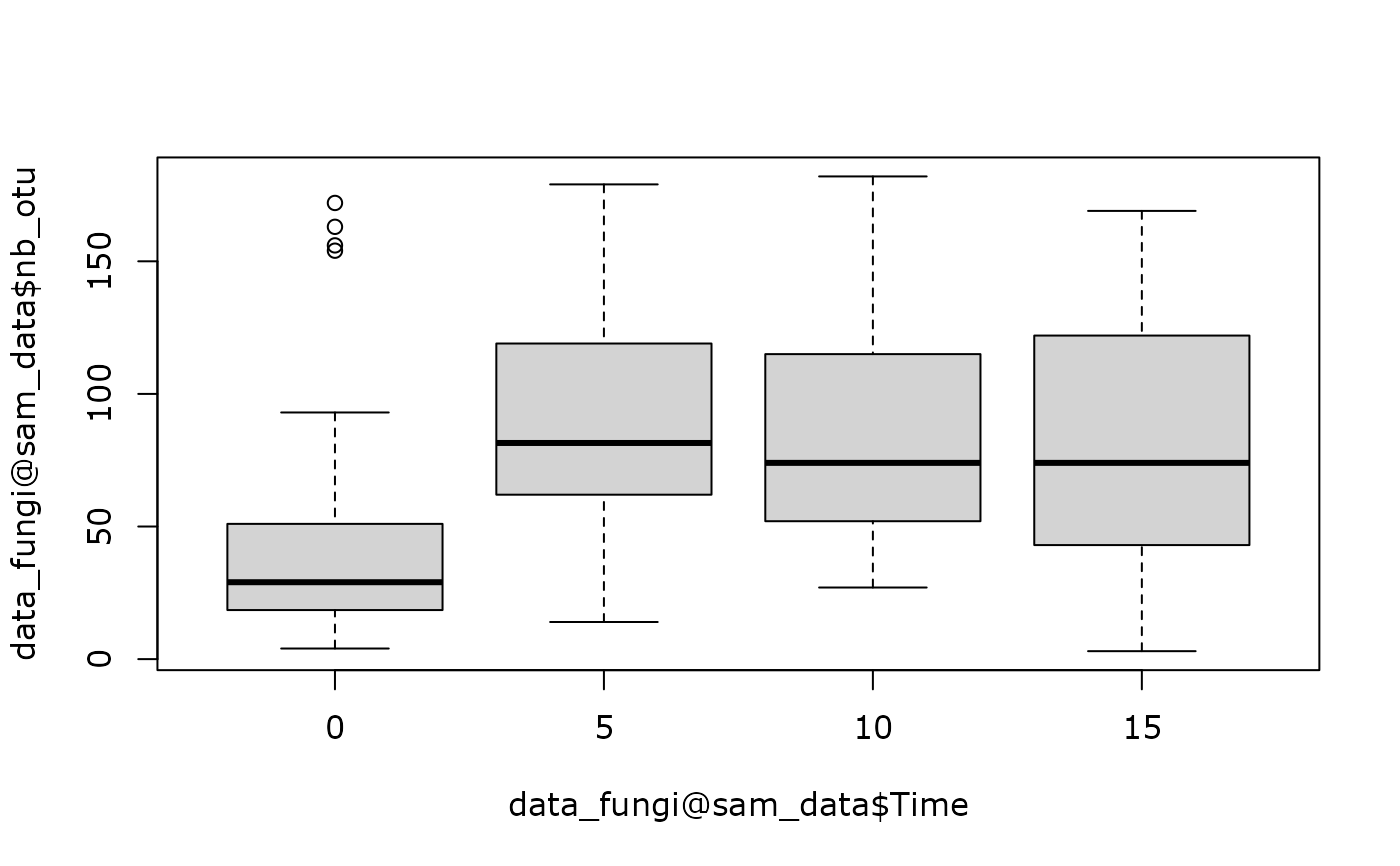
boxplot(data_fungi@sam_data$nb_otu ~ data_fungi@sam_data$Time)
 new_df <- data.frame(
variable_1 = runif(n = nsamples(data_fungi), min = 1, max = 20),
variable_2 = runif(n = nsamples(data_fungi), min = 1, max = 2)
)
rownames(new_df) <- sample_names(data_fungi)
data_fungi <- add_info_to_sam_data(data_fungi, new_df)
plot(data_fungi@sam_data$nb_otu ~ data_fungi@sam_data$variable_1)
new_df <- data.frame(
variable_1 = runif(n = nsamples(data_fungi), min = 1, max = 20),
variable_2 = runif(n = nsamples(data_fungi), min = 1, max = 2)
)
rownames(new_df) <- sample_names(data_fungi)
data_fungi <- add_info_to_sam_data(data_fungi, new_df)
plot(data_fungi@sam_data$nb_otu ~ data_fungi@sam_data$variable_1)