Calculate ecological distance among positive controls vs distance for all samples
Source:R/controls.R
dist_pos_control.RdCompute distance among positive controls, i.e. samples which are duplicated to test for variation, for example in (i) a step in the sampling, (ii) a step in the extraction, (iii) a step in the sequencing.
Arguments
- physeq
(required) a
phyloseq-classobject obtained using thephyloseqpackage.- samples_names
(required) a vector of names for samples with positives controls of the same samples having the same name
- method
(default: "bray") a method to calculate the distance, parsed to
vegan::vegdist(). See ?vegdist for a list of possible values.
Value
A list of two data-frames with (i) the distance among positive controls and (ii) the distance among all samples
Examples
data("enterotype")
sam_name_factice <- gsub("TS1_V2", "TS10_V2", sample_names(enterotype))
res_dist_cont <- dist_pos_control(enterotype, sam_name_factice)
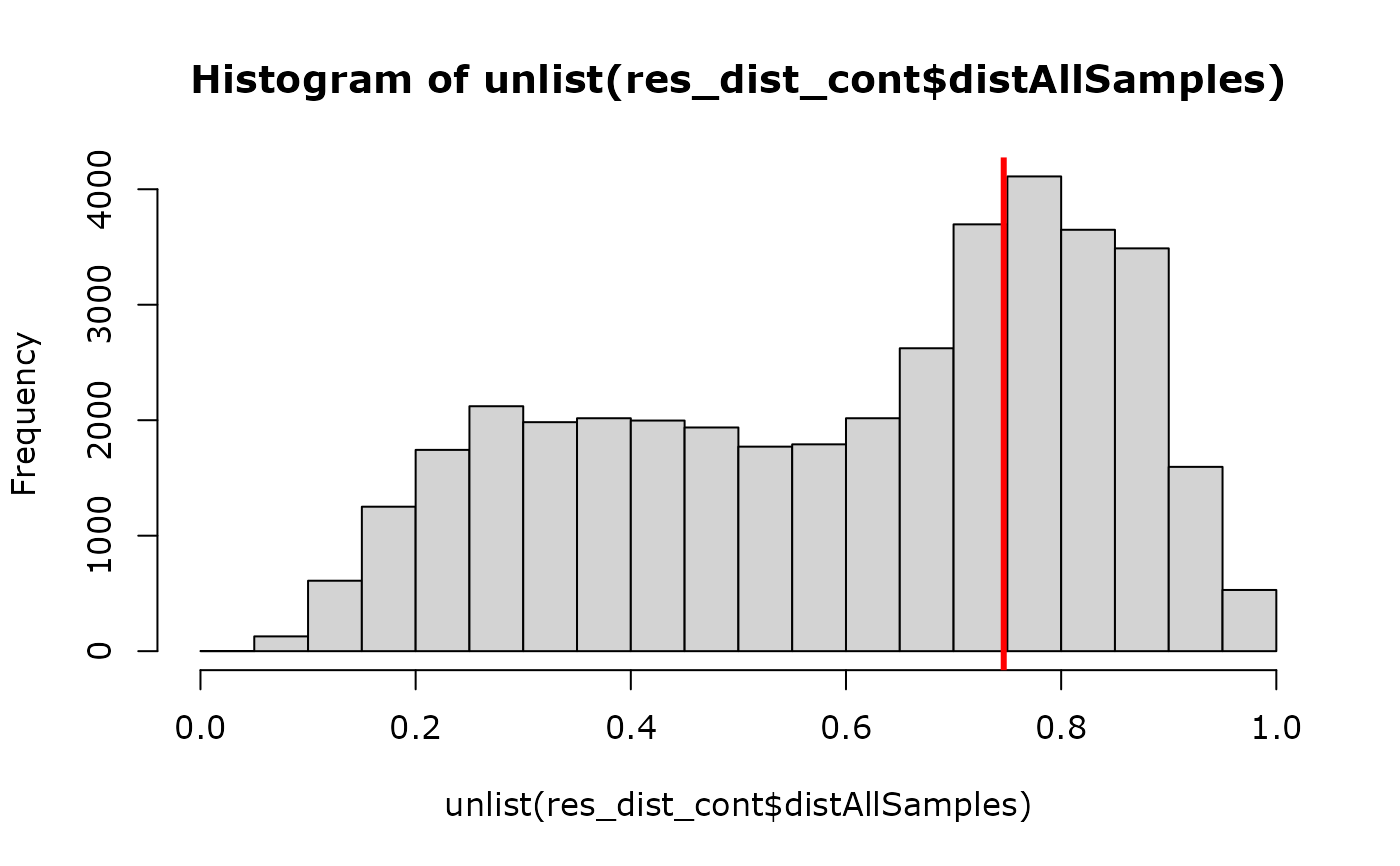
hist(unlist(res_dist_cont$distAllSamples))
abline(
v = mean(unlist(res_dist_cont$dist_controlontrolSamples), na.rm = TRUE),
col = "red", lwd = 3
)