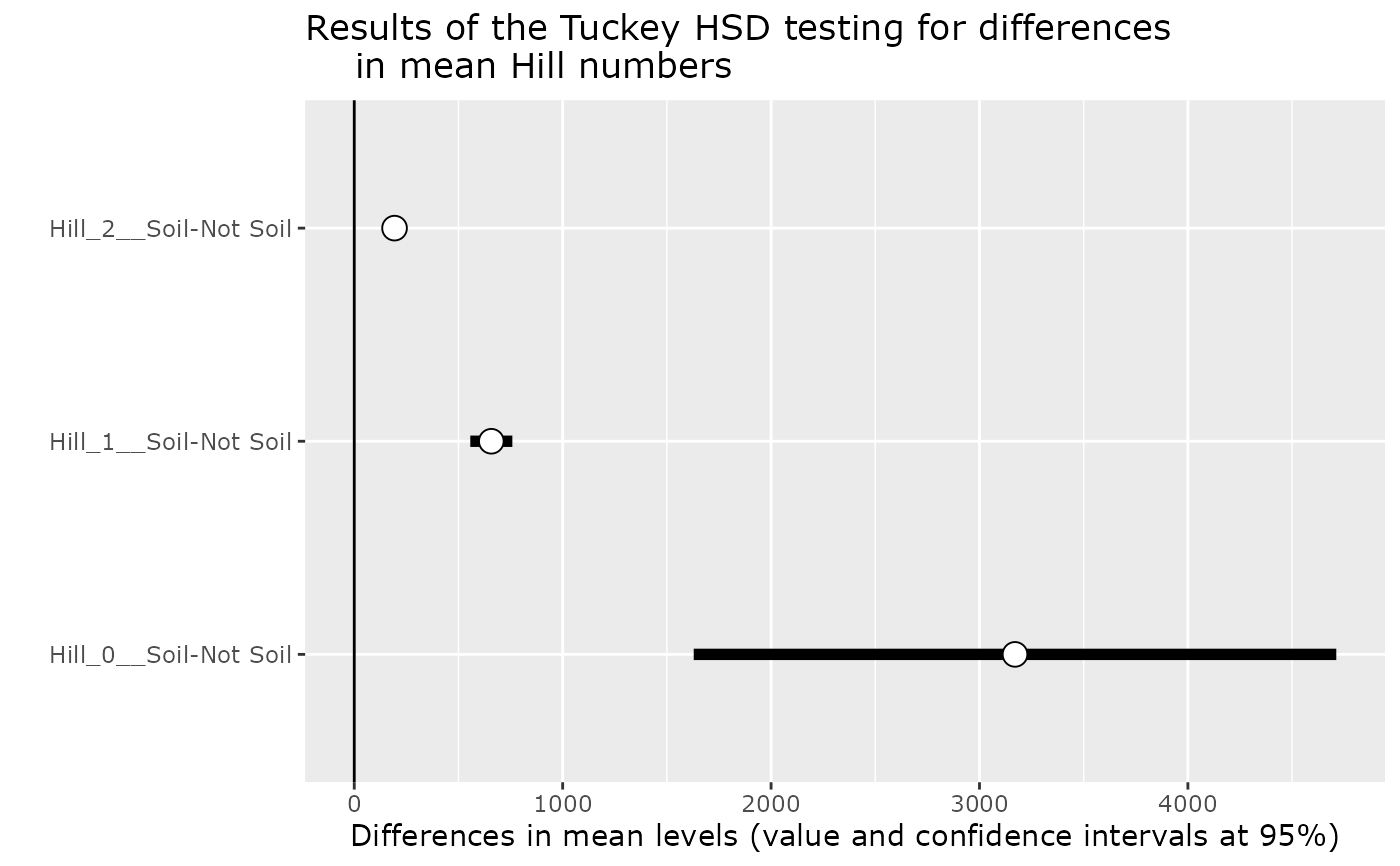
Note that, by default, this function use a sqrt of the read numbers in the linear model in order to correct for uneven sampling depth.
Usage
hill_tuckey_pq(
physeq,
modality,
q = c(0, 1, 2),
hill_scales = lifecycle::deprecated(),
silent = TRUE,
correction_for_sample_size = TRUE,
...
)Arguments
- physeq
(required) a
phyloseq-classobject obtained using thephyloseqpackage.- modality
(required) the variable to test
- q
(numeric vector) Hill diversity orders to compute (q values). Default computes Hill number 0 (species richness), Hill number 1 (exponential of Shannon index) and Hill number 2 (inverse of Simpson index). Formerly
hill_scales. Hill numbers are more appropriate in DNA metabarcoding studies whenq > 0(Alberdi & Gilbert, 2019; Calderón-Sanou et al., 2019).- hill_scales
- silent
(logical) If TRUE, no message are printing.
- correction_for_sample_size
(logical, default TRUE) This function use a sqrt of the read numbers in the linear model in order to correct for uneven sampling depth.
- ...
Additional arguments passed to
divent_hill_matrix_pq()and hence todivent::div_hill()(e.g.estimator = "naive"to match vegan-style results).
References
Alberdi, A., & Gilbert, M. T. P. (2019). A guide to the application of Hill numbers to DNA-based diversity analyses. Molecular Ecology Resources. doi:10.1111/1755-0998.13014
Calderón-Sanou, I., Münkemüller, T., Boyer, F., Zinger, L., & Thuiller, W. (2019). From environmental DNA sequences to ecological conclusions: How strong is the influence of methodological choices? Journal of Biogeography, 47. doi:10.1111/jbi.13681