A wrapper of plot_ordination with vegan distance matrix
Source:R/plot_functions.R
plot_ordination_pq.RdA wrapper of plot_ordination with vegan distance matrix
Arguments
- physeq
(required) a
phyloseq-classobject obtained using thephyloseqpackage.- method
(string, default "robust.aitchison") The distance method to use from vegan::vegdist(). See ?vegan::vegdist for more details.
- ordination_method
(string, default "NMDS") The ordination method to use in phyloseq::ordinate(). See ?phyloseq::ordinate for more details.
- ...
Additional arguments passed on to phyloseq::plot_ordination()
Details
Basically a wrapper of phyloseq::plot_ordination() to use aitchison and
robust.aitchison distances from vegan package.
Examples
library(patchwork)
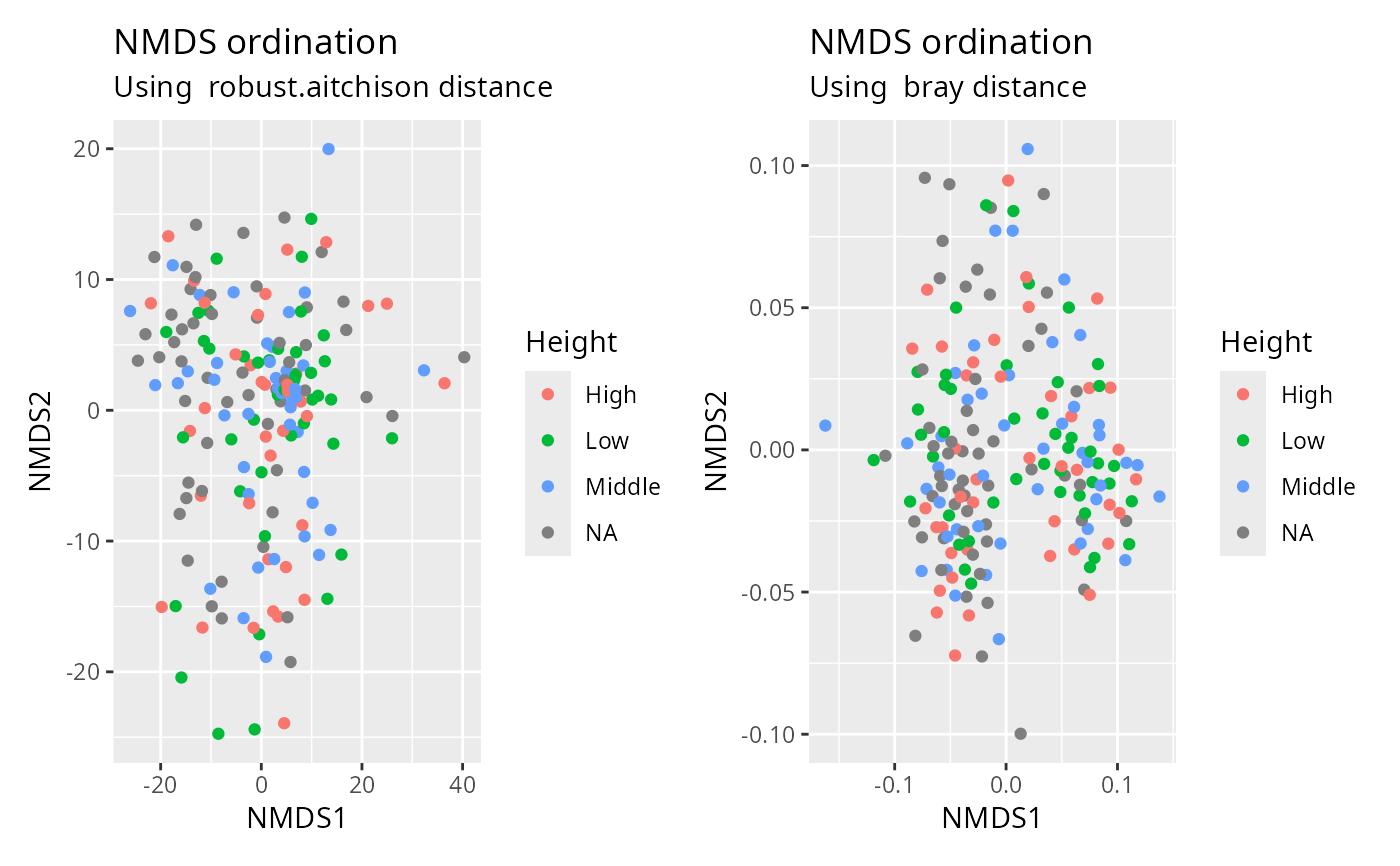
plot_ordination_pq(data_fungi_mini, method = "robust.aitchison", color = "Height") +
plot_ordination_pq(data_fungi_mini, method = "bray", color = "Height")
#> Taxa are now in columns.
#> Run 0 stress 0.115973
#> Run 1 stress 0.1300024
#> Run 2 stress 0.1183097
#> Run 3 stress 0.1214998
#> Run 4 stress 0.133196
#> Run 5 stress 0.1530583
#> Run 6 stress 0.1430454
#> Run 7 stress 0.1300024
#> Run 8 stress 0.1187081
#> Run 9 stress 0.1381296
#> Run 10 stress 0.1267024
#> Run 11 stress 0.1188736
#> Run 12 stress 0.1443663
#> Run 13 stress 0.1182422
#> Run 14 stress 0.118866
#> Run 15 stress 0.1188301
#> Run 16 stress 0.130007
#> Run 17 stress 0.1247712
#> Run 18 stress 0.1386751
#> Run 19 stress 0.1159623
#> ... New best solution
#> ... Procrustes: rmse 0.001485213 max resid 0.01705792
#> Run 20 stress 0.1381296
#> *** Best solution was not repeated -- monoMDS stopping criteria:
#> 13: stress ratio > sratmax
#> 7: scale factor of the gradient < sfgrmin
#> Warning: `aes_string()` was deprecated in ggplot2 3.0.0.
#> ℹ Please use tidy evaluation idioms with `aes()`.
#> ℹ See also `vignette("ggplot2-in-packages")` for more information.
#> ℹ The deprecated feature was likely used in the phyloseq package.
#> Please report the issue at <https://github.com/joey711/phyloseq/issues>.
#> Taxa are now in columns.
#> Run 0 stress 0.1894383
#> Run 1 stress 0.1888345
#> ... New best solution
#> ... Procrustes: rmse 0.0524328 max resid 0.2341065
#> Run 2 stress 0.1901206
#> Run 3 stress 0.1855488
#> ... New best solution
#> ... Procrustes: rmse 0.05964591 max resid 0.2629411
#> Run 4 stress 0.1884944
#> Run 5 stress 0.1886409
#> Run 6 stress 0.1865063
#> Run 7 stress 0.1873102
#> Run 8 stress 0.190706
#> Run 9 stress 0.1873528
#> Run 10 stress 0.1882318
#> Run 11 stress 0.189307
#> Run 12 stress 0.1896571
#> Run 13 stress 0.1864311
#> Run 14 stress 0.1893721
#> Run 15 stress 0.1854113
#> ... New best solution
#> ... Procrustes: rmse 0.01940789 max resid 0.1180019
#> Run 16 stress 0.1904496
#> Run 17 stress 0.1869919
#> Run 18 stress 0.1887375
#> Run 19 stress 0.1902213
#> Run 20 stress 0.1902424
#> *** Best solution was not repeated -- monoMDS stopping criteria:
#> 14: no. of iterations >= maxit
#> 6: stress ratio > sratmax