Computes a manifold approximation and projection (UMAP) for phyloseq object
Source:R/plot_functions.R
umap_pq.RdArguments
- physeq
(required) a
phyloseq-classobject obtained using thephyloseqpackage.- pkg
Which R packages to use, either "umap" or "uwot".
- ...
Additional arguments passed on to
umap::umap()oruwot::umap2()function. For examplen_neighborsset the number of nearest neighbors (Default 15). Seeumap::umap.defaults()oruwot::umap2()for the list of parameters and default values.
Details
This function is mainly a wrapper of the work of others.
Please make a reference to umap::umap() if you
use this function.
Examples
library("umap")
df_umap <- umap_pq(data_fungi_mini)
#> Taxa are now in columns.
#> Taxa are now in rows.
#> Joining with `by = join_by(Sample)`
#> Warning: The `x` argument of `as_tibble.matrix()` must have unique column names if
#> `.name_repair` is omitted as of tibble 2.0.0.
#> ℹ Using compatibility `.name_repair`.
#> ℹ The deprecated feature was likely used in the MiscMetabar package.
#> Please report the issue at
#> <https://github.com/adrientaudiere/MiscMetabar/issues>.
#> Joining with `by = join_by(Sample)`
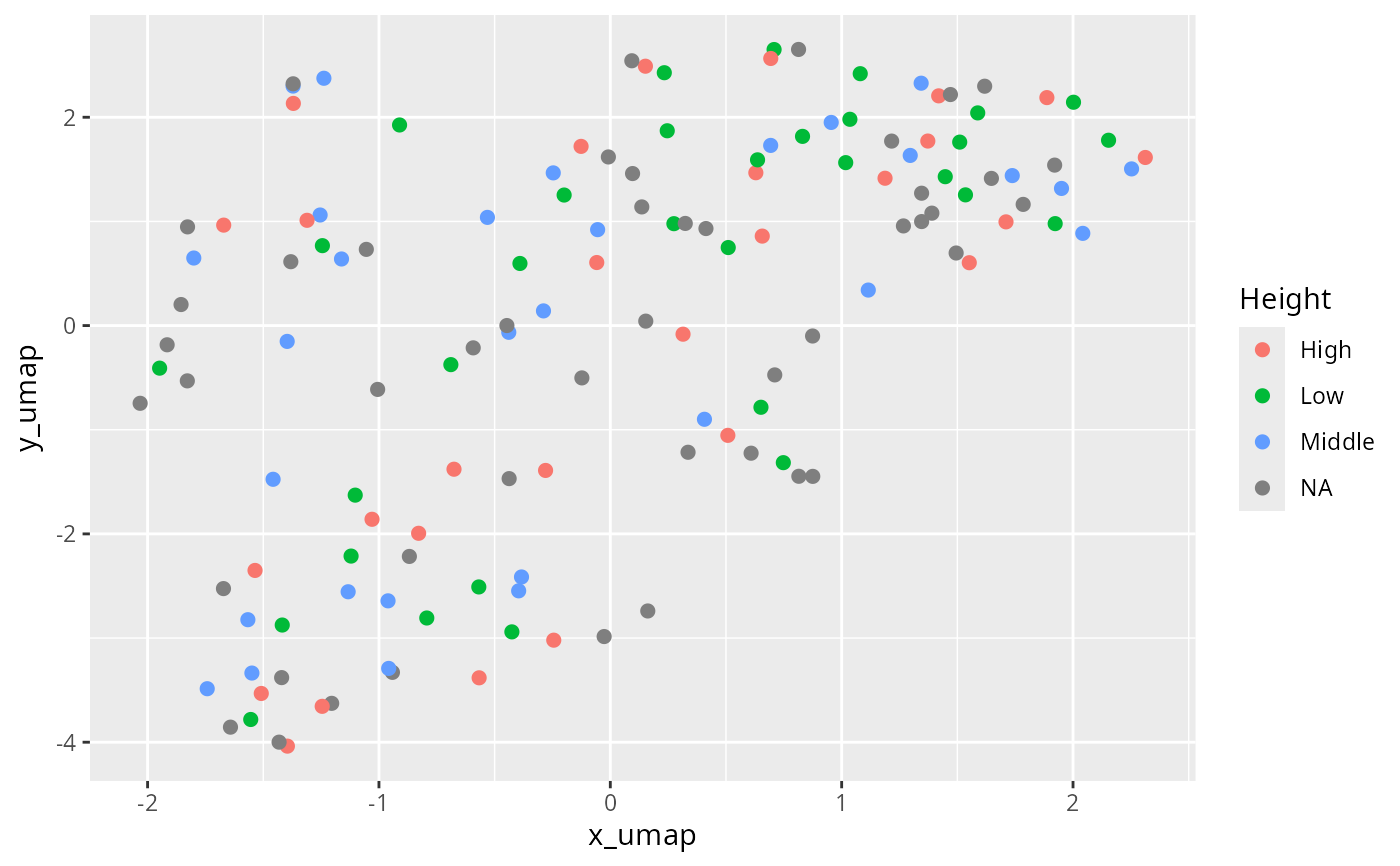
ggplot(df_umap, aes(x = x_umap, y = y_umap, col = Height)) +
geom_point(size = 2)
 # \donttest{
library(patchwork)
physeq <- data_fungi_mini
res_tsne <- tsne_pq(data_fungi_mini)
df_umap_tsne <- df_umap
df_umap_tsne$x_tsne <- res_tsne$Y[, 1]
df_umap_tsne$y_tsne <- res_tsne$Y[, 2]
((ggplot(df_umap, aes(x = x_umap, y = y_umap, col = Height)) +
geom_point(size = 2) +
ggtitle("UMAP")) + (plot_ordination(physeq,
ordination =
ordinate(physeq, method = "PCoA", distance = "bray"), color = "Height"
)) +
ggtitle("PCoA")) /
((ggplot(df_umap_tsne, aes(x = x_tsne, y = y_tsne, col = Height)) +
geom_point(size = 2) +
ggtitle("tsne")) +
(plot_ordination(physeq,
ordination = ordinate(physeq, method = "NMDS", distance = "bray"),
color = "Height"
) +
ggtitle("NMDS"))) +
patchwork::plot_layout(guides = "collect")
#> Square root transformation
#> Wisconsin double standardization
#> Run 0 stress 0.1842417
#> Run 1 stress 0.1830653
#> ... New best solution
#> ... Procrustes: rmse 0.05975443 max resid 0.2477283
#> Run 2 stress 0.1808269
#> ... New best solution
#> ... Procrustes: rmse 0.07519717 max resid 0.2637364
#> Run 3 stress 0.1802656
#> ... New best solution
#> ... Procrustes: rmse 0.04975387 max resid 0.2982434
#> Run 4 stress 0.183484
#> Run 5 stress 0.184756
#> Run 6 stress 0.1847681
#> Run 7 stress 0.1819892
#> Run 8 stress 0.1820539
#> Run 9 stress 0.18634
#> Run 10 stress 0.1856027
#> Run 11 stress 0.1827823
#> Run 12 stress 0.1816633
#> Run 13 stress 0.1833599
#> Run 14 stress 0.1842722
#> Run 15 stress 0.1850547
#> Run 16 stress 0.1860664
#> Run 17 stress 0.1857189
#> Run 18 stress 0.1795345
#> ... New best solution
#> ... Procrustes: rmse 0.05055477 max resid 0.2998752
#> Run 19 stress 0.1857847
#> Run 20 stress 0.1831573
#> *** Best solution was not repeated -- monoMDS stopping criteria:
#> 18: no. of iterations >= maxit
#> 2: stress ratio > sratmax
# \donttest{
library(patchwork)
physeq <- data_fungi_mini
res_tsne <- tsne_pq(data_fungi_mini)
df_umap_tsne <- df_umap
df_umap_tsne$x_tsne <- res_tsne$Y[, 1]
df_umap_tsne$y_tsne <- res_tsne$Y[, 2]
((ggplot(df_umap, aes(x = x_umap, y = y_umap, col = Height)) +
geom_point(size = 2) +
ggtitle("UMAP")) + (plot_ordination(physeq,
ordination =
ordinate(physeq, method = "PCoA", distance = "bray"), color = "Height"
)) +
ggtitle("PCoA")) /
((ggplot(df_umap_tsne, aes(x = x_tsne, y = y_tsne, col = Height)) +
geom_point(size = 2) +
ggtitle("tsne")) +
(plot_ordination(physeq,
ordination = ordinate(physeq, method = "NMDS", distance = "bray"),
color = "Height"
) +
ggtitle("NMDS"))) +
patchwork::plot_layout(guides = "collect")
#> Square root transformation
#> Wisconsin double standardization
#> Run 0 stress 0.1842417
#> Run 1 stress 0.1830653
#> ... New best solution
#> ... Procrustes: rmse 0.05975443 max resid 0.2477283
#> Run 2 stress 0.1808269
#> ... New best solution
#> ... Procrustes: rmse 0.07519717 max resid 0.2637364
#> Run 3 stress 0.1802656
#> ... New best solution
#> ... Procrustes: rmse 0.04975387 max resid 0.2982434
#> Run 4 stress 0.183484
#> Run 5 stress 0.184756
#> Run 6 stress 0.1847681
#> Run 7 stress 0.1819892
#> Run 8 stress 0.1820539
#> Run 9 stress 0.18634
#> Run 10 stress 0.1856027
#> Run 11 stress 0.1827823
#> Run 12 stress 0.1816633
#> Run 13 stress 0.1833599
#> Run 14 stress 0.1842722
#> Run 15 stress 0.1850547
#> Run 16 stress 0.1860664
#> Run 17 stress 0.1857189
#> Run 18 stress 0.1795345
#> ... New best solution
#> ... Procrustes: rmse 0.05055477 max resid 0.2998752
#> Run 19 stress 0.1857847
#> Run 20 stress 0.1831573
#> *** Best solution was not repeated -- monoMDS stopping criteria:
#> 18: no. of iterations >= maxit
#> 2: stress ratio > sratmax
 df_uwot <- umap_pq(data_fungi_mini, pkg = "uwot")
#> Taxa are now in columns.
#> Taxa are now in rows.
#> Joining with `by = join_by(Sample)`
#> Joining with `by = join_by(Sample)`
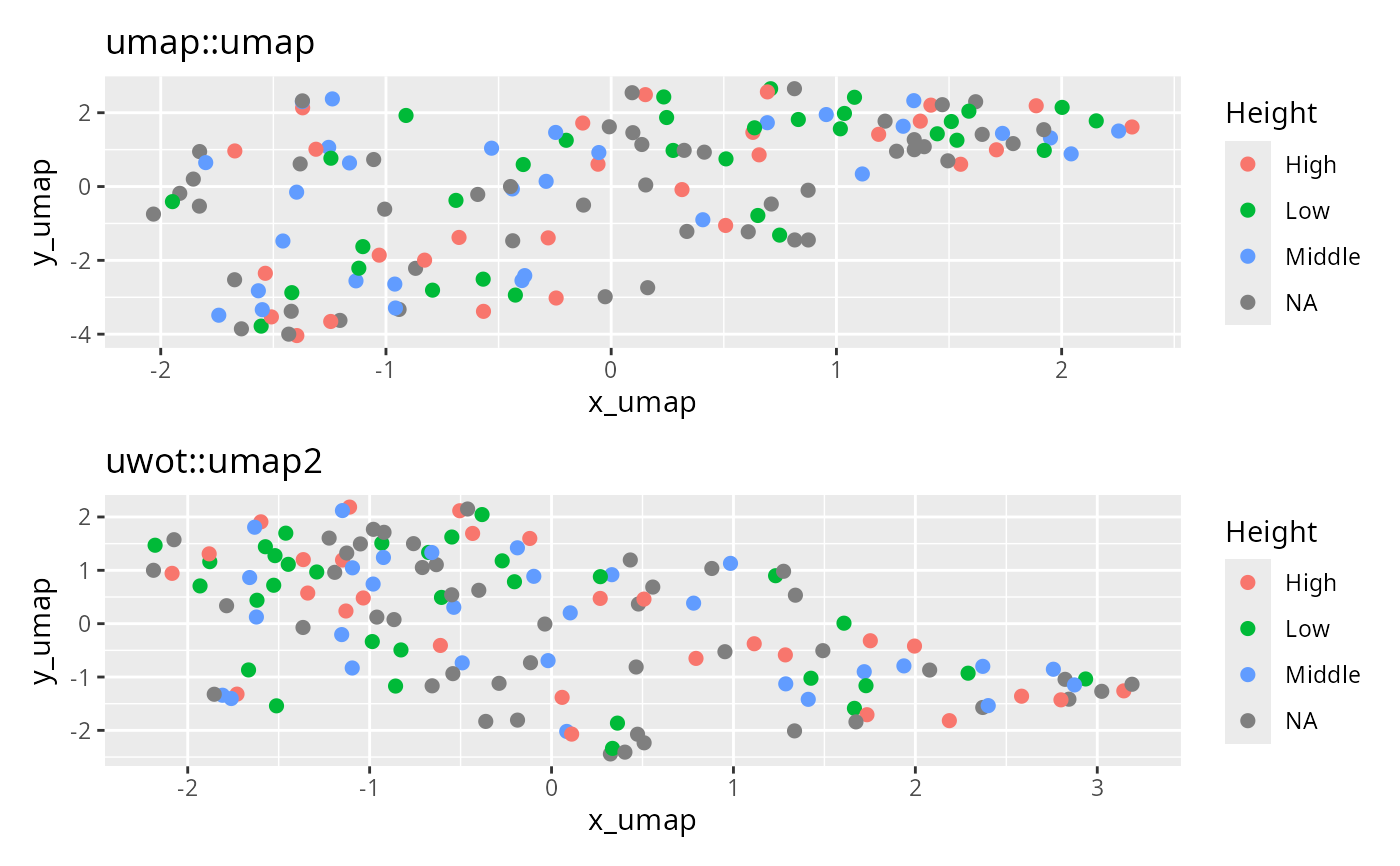
(ggplot(df_umap, aes(x = x_umap, y = y_umap, col = Height)) +
geom_point(size = 2) +
ggtitle("umap::umap")) /
(ggplot(df_uwot, aes(x = x_umap, y = y_umap, col = Height)) +
geom_point(size = 2) +
ggtitle("uwot::umap2"))
df_uwot <- umap_pq(data_fungi_mini, pkg = "uwot")
#> Taxa are now in columns.
#> Taxa are now in rows.
#> Joining with `by = join_by(Sample)`
#> Joining with `by = join_by(Sample)`
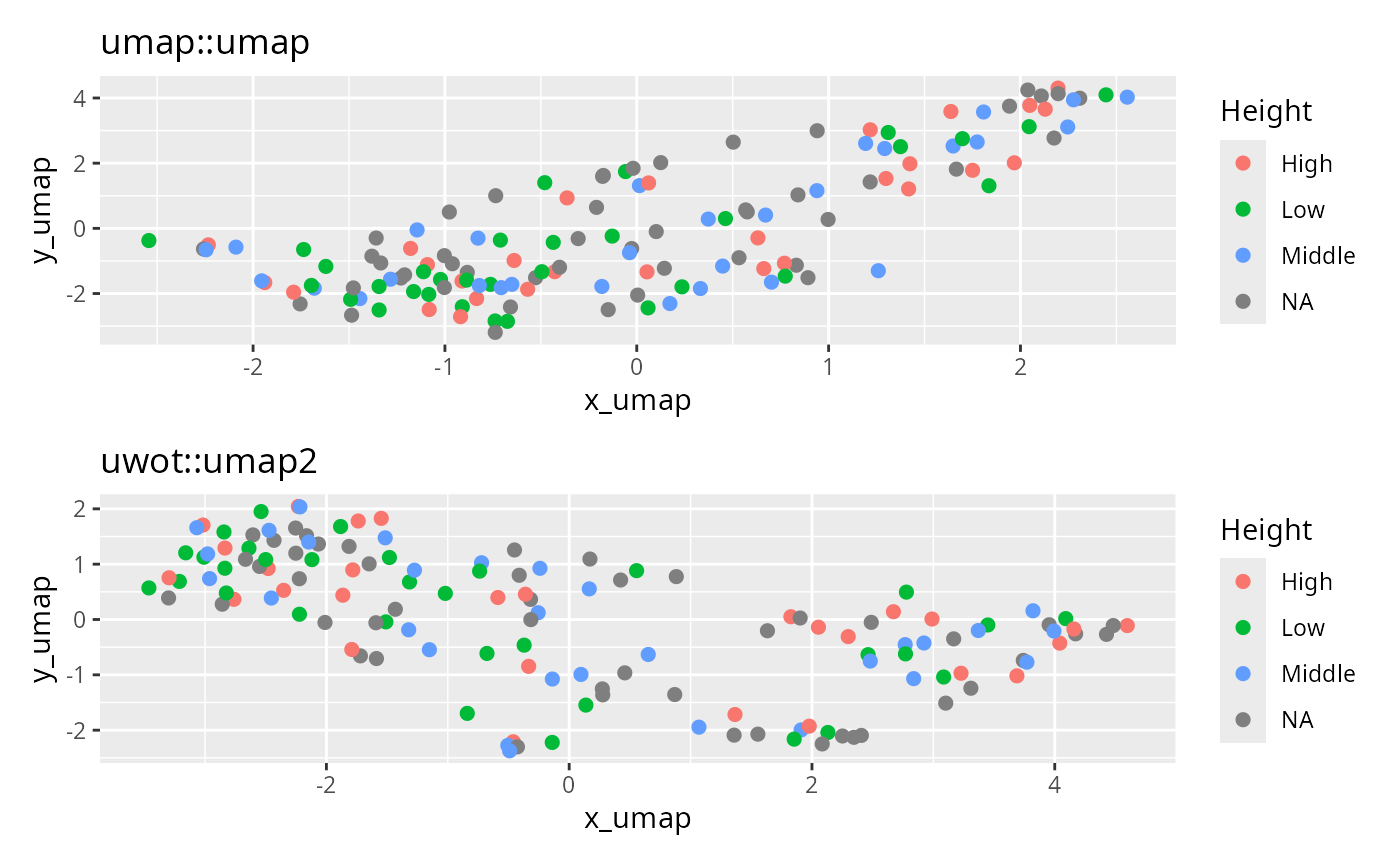
(ggplot(df_umap, aes(x = x_umap, y = y_umap, col = Height)) +
geom_point(size = 2) +
ggtitle("umap::umap")) /
(ggplot(df_uwot, aes(x = x_umap, y = y_umap, col = Height)) +
geom_point(size = 2) +
ggtitle("uwot::umap2"))
 # }
# }
