Unified dispatcher for all OTU-table transformations and normalisations
Source:R/normalize_pq.R
transform_pq.RdSingle entry-point for all count-table transformations available in
MiscMetabar. Ecological methods ("tss", "hellinger", "clr",
"rclr", "log1p", "z", "pa", "rank") are delegated to
vegan::decostand(). Library-size normalisation methods ("rarefy",
"srs", "gmpr", "css", "tmm", "vst") and the McKnight
log-log residual method ("mcknight_residuals") are delegated to
their dedicated *_pq() functions. All ... arguments are forwarded
to the underlying function.
Usage
transform_pq(
physeq,
method = c("tss", "hellinger", "clr", "rclr", "log1p", "z", "pa", "rank",
"normalize_prop", "rarefy", "srs", "gmpr", "css", "tmm", "vst", "mcknight_residuals"),
pseudocount = 1,
...
)Arguments
- physeq
(required) a
phyloseq-classobject obtained using thephyloseqpackage.- method
(character, default
"tss") transformation to apply. One of:"tss"Total Sum Scaling — divide by library size.
"hellinger"Square-root of proportions. Good for ordination.
"clr"Centred log-ratio (adds
pseudocountto handle zeros)."rclr"Robust CLR (adds
pseudocountto handle zeros)."log1p"\(\log(1 + x)\) transformation.
"z"Per-taxon z-score standardisation.
"pa"Presence/absence (0/1).
"rank"Replace counts by within-sample ranks.
"normalize_prop"TSS × constant + log, via
normalize_prop_pq()."rarefy"Rarefaction to equal depth, via
rarefy_pq()."srs"Scaling with Ranked Subsampling, via
srs_pq(). Requires the SRS package."gmpr"Geometric Mean of Pairwise Ratios, via
gmpr_pq()."css"Cumulative Sum Scaling, via
css_pq(). Requires the metagenomeSeq package."tmm"Trimmed Mean of M-values, via
tmm_pq(). Requires the edgeR package."vst"Variance Stabilising Transformation, via
vst_pq(). Requires the DESeq2 package."mcknight_residuals"Log-log depth residuals added to
sample_data, viamcknight_residuals_pq().
- pseudocount
(numeric, default
1) added before"clr"/"rclr"to avoid non-positive values. Ignored for all other methods.- ...
Additional arguments forwarded to the underlying function (
vegan::decostand(),rarefy_pq(),srs_pq(), etc.).
Value
A new phyloseq-class object with a
transformed otu_table (and augmented sample_data for
"mcknight_residuals").
Examples
data_f_tss <- transform_pq(data_fungi_mini, method = "tss")
sample_sums(data_f_tss)
#> A10-005-B_S188_MERGED.fastq.gz A10-005-H_S189_MERGED.fastq.gz
#> 1 1
#> A10-005-M_S190_MERGED.fastq.gz A12-007_S191_MERGED.fastq.gz
#> 1 1
#> A12-007-B_S2_MERGED.fastq.gz A15-004_S3_MERGED.fastq.gz
#> 1 1
#> A8-005_S4_MERGED.fastq.gz AB29-ABMX-H_S6_MERGED.fastq.gz
#> 1 1
#> AC27-013_S7_MERGED.fastq.gz AC29033_S8_MERGED.fastq.gz
#> 1 1
#> AD26-005-B_S9_MERGED.fastq.gz AD26-005-H_S10_MERGED.fastq.gz
#> 1 1
#> AD26-005-M_S11_MERGED.fastq.gz AD30-ABMX-M_S12_MERGED.fastq.gz
#> 1 1
#> AD32-007-M_S13_MERGED.fastq.gz ADABM30X-B_S14_MERGED.fastq.gz
#> 1 1
#> ADABM30X-H_S15_MERGED.fastq.gz ADABM30X-M_S16_MERGED.fastq.gz
#> 1 1
#> AE30-ABM507_S17_MERGED.fastq.gz B17-014_S18_MERGED.fastq.gz
#> 1 1
#> B18-006-B_S19_MERGED.fastq.gz BA16-036bis_S20_MERGED.fastq.gz
#> 1 1
#> BA17-050-B_S21_MERGED.fastq.gz BB19-006-H_S22_MERGED.fastq.gz
#> 1 1
#> BB6-019-M_S25_MERGED.fastq.gz BD14-021_S26_MERGED.fastq.gz
#> 1 1
#> BG7-010-H_S31_MERGED.fastq.gz BH9-021_S33_MERGED.fastq.gz
#> 1 1
#> BJ17-007-M_S34_MERGED.fastq.gz BJ8-ABM-003_S35_MERGED.fastq.gz
#> 1 1
#> BL7-006-H_S37_MERGED.fastq.gz BN11-041_S39_MERGED.fastq.gz
#> 1 1
#> BO8-002_S41_MERGED.fastq.gz BO8-005_S42_MERGED.fastq.gz
#> 1 1
#> BP11-001-B_S43_MERGED.fastq.gz BP11-001-H_S44_MERGED.fastq.gz
#> 1 1
#> BP11-001-M_S45_MERGED.fastq.gz BP12-025-B_S46_MERGED.fastq.gz
#> 1 1
#> BP14-006_S47_MERGED.fastq.gz BQ3-019_S48_MERGED.fastq.gz
#> 1 1
#> BQ4-018-B_S49_MERGED.fastq.gz BQ4-018-H_S50_MERGED.fastq.gz
#> 1 1
#> BQ4-018-M_S51_MERGED.fastq.gz BQ9ABM-002_S52_MERGED.fastq.gz
#> 1 1
#> BR8-005_S53_MERGED.fastq.gz BS14-006_S54_MERGED.fastq.gz
#> 1 1
#> BT-006-M_S55_MERGED.fastq.gz BT7-006_S56_MERGED.fastq.gz
#> 1 1
#> BV11-002-B_S57_MERGED.fastq.gz BV11-002-H_S58_MERGED.fastq.gz
#> 1 1
#> BV11-002-M_S59_MERGED.fastq.gz BW8-003_S60_MERGED.fastq.gz
#> 1 1
#> C1-001_S61_MERGED.fastq.gz C21-NV1-B_S62_MERGED.fastq.gz
#> 1 1
#> C9-005_S65_MERGED.fastq.gz CA9-X_S68_MERGED.fastq.gz
#> 1 1
#> CB8-019-B_S69_MERGED.fastq.gz CB8-019-H_S70_MERGED.fastq.gz
#> 1 1
#> CB8-019-M_S71_MERGED.fastq.gz CB9-013_S72_MERGED.fastq.gz
#> 1 1
#> CC3-044_S73_MERGED.fastq.gz D17-011_S77_MERGED.fastq.gz
#> 1 1
#> D18-003-B_S78_MERGED.fastq.gz D18-003-M_S80_MERGED.fastq.gz
#> 1 1
#> D22-NVABM1_S81_MERGED.fastq.gz D61-010-B_S82_MERGED.fastq.gz
#> 1 1
#> D9-027-H_S84_MERGED.fastq.gz D9-027-M_S85_MERGED.fastq.gz
#> 1 1
#> DBM-ABM-001_S86_MERGED.fastq.gz DJ2-008-H_S88_MERGED.fastq.gz
#> 1 1
#> DP4-ABM001_S90_MERGED.fastq.gz DS1-ABM002-B_S91_MERGED.fastq.gz
#> 1 1
#> DS1-ABM002-H_S92_MERGED.fastq.gz DS1-ABM002-M_S93_MERGED.fastq.gz
#> 1 1
#> DU3-045-B_S94_MERGED.fastq.gz DW4-007_S95_MERGED.fastq.gz
#> 1 1
#> DY5-004-B_S96_MERGED.fastq.gz DY5-004-H_S97_MERGED.fastq.gz
#> 1 1
#> DZ6-ABM-001_S99_MERGED.fastq.gz EA5-ABM-001_S103_MERGED.fastq.gz
#> 1 1
#> EC2-013-B_S104_MERGED.fastq.gz F6-ABM-001_S105_MERGED.fastq.gz
#> 1 1
#> F7-015-M_S106_MERGED.fastq.gz FOMES19-H_S108_MERGED.fastq.gz
#> 1 1
#> FOMES19-M_S109_MERGED.fastq.gz H10-018-M_S110_MERGED.fastq.gz
#> 1 1
#> H24-NVABM1-H_S111_MERGED.fastq.gz J18-004-B_S114_MERGED.fastq.gz
#> 1 1
#> J18-004-M_S116_MERGED.fastq.gz K18-002-H_S117_MERGED.fastq.gz
#> 1 1
#> K26-NVABM1_S118_MERGED.fastq.gz L19X-B_S119_MERGED.fastq.gz
#> 1 1
#> L19X-H_S120_MERGED.fastq.gz L19X-M_S121_MERGED.fastq.gz
#> 1 1
#> L23-002-B_S122_MERGED.fastq.gz L23-002-H_S123_MERGED.fastq.gz
#> 1 1
#> L23-002-M_S124_MERGED.fastq.gz N19X-H_S127_MERGED.fastq.gz
#> 1 1
#> N19X-M_S128_MERGED.fastq.gz N23-002-B_S130_MERGED.fastq.gz
#> 1 1
#> N23-002-H_S131_MERGED.fastq.gz N25-ABMX_S133_MERGED.fastq.gz
#> 1 1
#> NVABM-0058_S134_MERGED.fastq.gz NVABM-0163-H_S135_MERGED.fastq.gz
#> 1 1
#> NVABM-0397_S138_MERGED.fastq.gz NVABM0216_S136_MERGED.fastq.gz
#> 1 1
#> NVABM0244-M_S137_MERGED.fastq.gz O20-X-H_S140_MERGED.fastq.gz
#> 1 1
#> O20-X-M_S141_MERGED.fastq.gz O24-003-B_S145_MERGED.fastq.gz
#> 1 1
#> O26-004-B_S148_MERGED.fastq.gz O27-012_S151_MERGED.fastq.gz
#> 1 1
#> O9-005-B_S152_MERGED.fastq.gz P19-023-M_S153_MERGED.fastq.gz
#> 1 1
#> P27-015-M_S154_MERGED.fastq.gz P27-ABM001_S155_MERGED.fastq.gz
#> 1 1
#> Q27-ABM003-B_S156_MERGED.fastq.gz R25-ABMX_S157_MERGED.fastq.gz
#> 1 1
#> T28-011_S161_MERGED.fastq.gz T28-ABM602-B_S162_MERGED.fastq.gz
#> 1 1
#> U27-ABM002_S163_MERGED.fastq.gz W25-ABMX_S164_MERGED.fastq.gz
#> 1 1
#> W26-001-B_S165_MERGED.fastq.gz W30-006_S168_MERGED.fastq.gz
#> 1 1
#> W9-025-M_S169_MERGED.fastq.gz X24-009-B_S170_MERGED.fastq.gz
#> 1 1
#> X24-009-H_S171_MERGED.fastq.gz X24-009-M_S172_MERGED.fastq.gz
#> 1 1
#> X24-010_S173_MERGED.fastq.gz X29-004-B_S174_MERGED.fastq.gz
#> 1 1
#> X29-004-H_S175_MERGED.fastq.gz X29-004-M_S176_MERGED.fastq.gz
#> 1 1
#> Y21-ABM484-H_S177_MERGED.fastq.gz Y28-002-B_S178_MERGED.fastq.gz
#> 1 1
#> Y31-ABM484-B_S184_MERGED.fastq.gz Z29-001-H_S185_MERGED.fastq.gz
#> 1 1
#> Z30-ABM560-M_S187_MERGED.fastq.gz
#> 1
# \donttest{
data_f_hell <- transform_pq(data_fungi_mini, method = "hellinger")
data_f_clr <- transform_pq(data_fungi_mini, method = "clr")
data_f_rclr <- transform_pq(data_fungi_mini, method = "rclr")
data_f_log1p <- transform_pq(data_fungi_mini, method = "log1p")
data_f_z <- transform_pq(data_fungi_mini, method = "z")
data_f_pa <- transform_pq(data_fungi_mini, method = "pa")
data_f_rank <- transform_pq(data_fungi_mini, method = "rank")
data_f_norm_prop_log10 <- transform_pq(data_fungi_mini,
method = "normalize_prop", base_log = 10)
data_f_norm_prop_no_log <- transform_pq(data_fungi_mini,
method = "normalize_prop", base_log = NULL)
data_f_norm_prop_log2 <- transform_pq(data_fungi_mini,
method = "normalize_prop", base_log = 2)
data_f_rarefy <- transform_pq(data_fungi_mini, method = "rarefy", seed = 1)
data_f_srs <- transform_pq(data_fungi_mini, method = "srs", seed = 1)
data_f_gmpr <- transform_pq(data_fungi_mini, method = "gmpr")
#> Warning: GMPR size factors could not be computed for 90 sample(s); these samples are left unscaled.
data_f_css <- transform_pq(data_fungi_mini, method = "css")
#> Warning: cumNormStatFast() failed (likely samples with <=1 feature); falling back to cumNormStat(). Original error: Warning sample with one or zero features
#> Default value being used.
data_f_tmm <- transform_pq(data_fungi_mini, method = "tmm")
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
#> Warning: no non-missing arguments to max; returning -Inf
data_f_vst <- transform_pq(data_fungi_mini, method = "vst")
#> converting counts to integer mode
#> -- note: fitType='parametric', but the dispersion trend was not well captured by the
#> function: y = a/x + b, and a local regression fit was automatically substituted.
#> specify fitType='local' or 'mean' to avoid this message next time.
data_f_mcknight <- transform_pq(data_fungi_mini, method = "mcknight_residuals")
otu_list <- list(
hell = unclass(data_f_hell@otu_table),
clr = unclass(data_f_clr@otu_table),
rclr = unclass(data_f_rclr@otu_table),
log1p = unclass(data_f_log1p@otu_table),
z = unclass(data_f_z@otu_table),
rarefy = unclass(data_f_rarefy@otu_table)
)
pairs_cor <- sapply(
otu_list,
\(x) sapply(otu_list, \(y) cor(as.vector(x), as.vector(y)))
)
pairs_cor
#> hell clr rclr log1p z rarefy
#> hell 1.0000000 0.8138757 0.8138757 0.8076421 0.5742337 0.7663504
#> clr 0.8138757 1.0000000 1.0000000 0.9809713 0.6929287 0.5807582
#> rclr 0.8138757 1.0000000 1.0000000 0.9809713 0.6929287 0.5807582
#> log1p 0.8076421 0.9809713 0.9809713 1.0000000 0.7028648 0.5697071
#> z 0.5742337 0.6929287 0.6929287 0.7028648 1.0000000 0.4800643
#> rarefy 0.7663504 0.5807582 0.5807582 0.5697071 0.4800643 1.0000000
plot(unclass(data_f_mcknight@otu_table), unclass(data_f_css@otu_table))
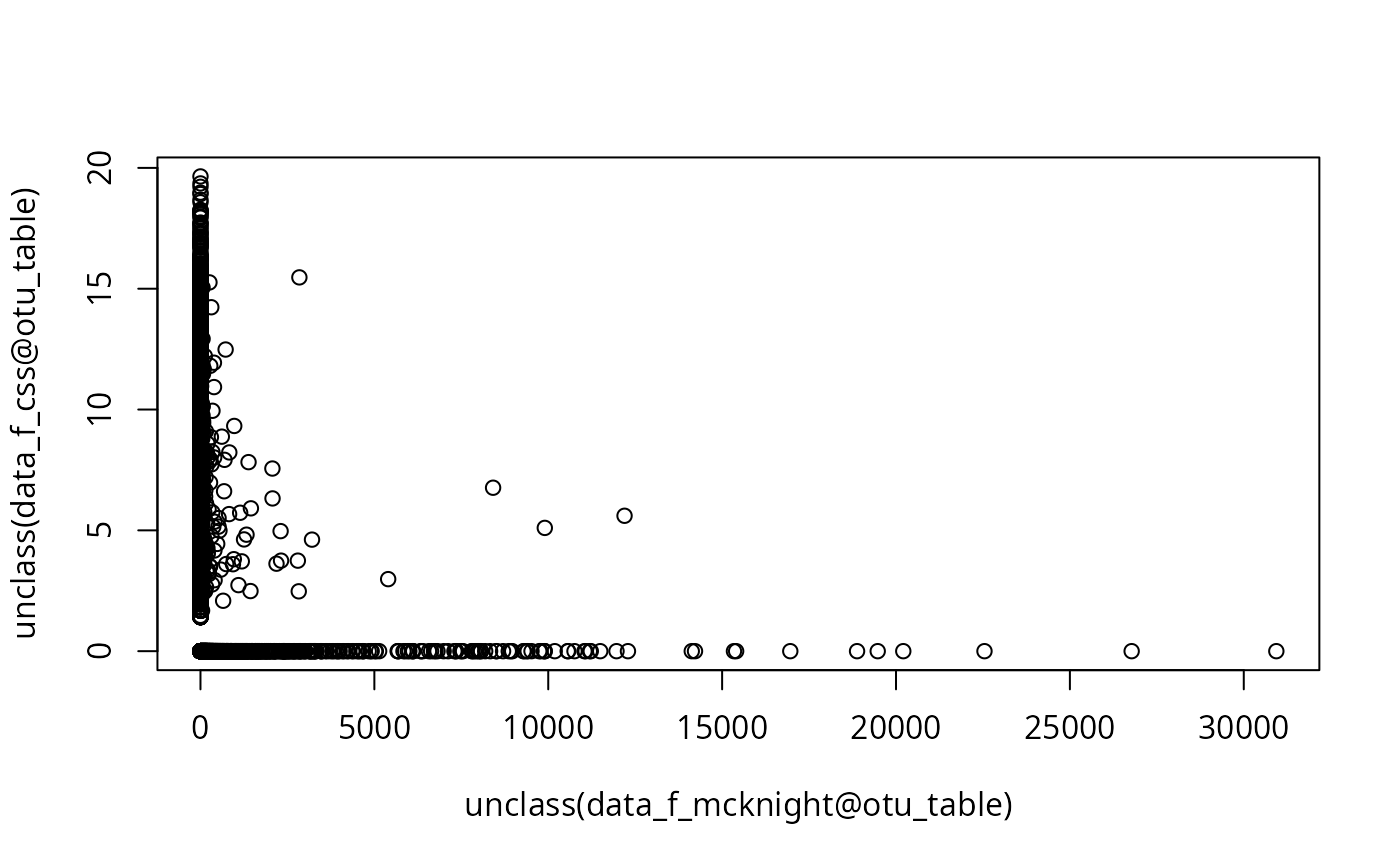 plot(unclass(data_f_rarefy@otu_table), unclass(data_f_clr@otu_table))
plot(unclass(data_f_rarefy@otu_table), unclass(data_f_clr@otu_table))
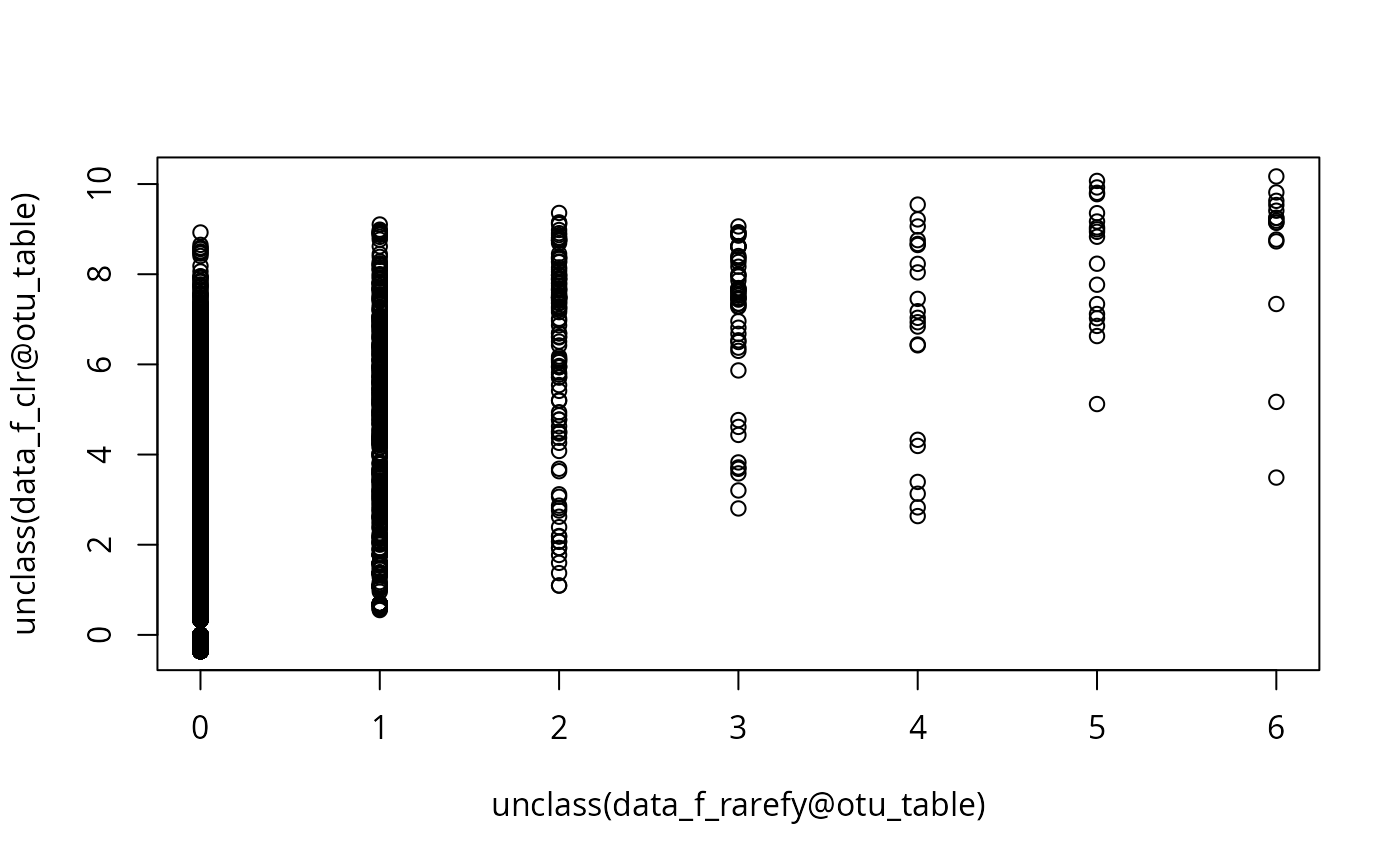 # }
# }
